The Ackermann function is a well-known example of a recursive function that is not primitive recursive. It serves as a benchmark for computing and illustrates the concept of deep recursion.
"Tube sound" refers to the characteristic audio quality produced by vacuum tube amplifiers, widely used in music production and amplification, particularly in electric guitars and high-fidelity audio systems. Tube amplifiers use vacuum tubes (also known as thermionic valves) to amplify audio signals, and they are known for creating a warm, rich, and pleasing sound.
EPNdB stands for "Equivalent Noise Power in Decibels." It is a metric used to quantify the noise performance of communication systems, particularly in the context of radio and telecommunications. EPNdB expresses the noise figure or noise performance in decibels, allowing for easier comparison of different systems or components. Essentially, EPNdB helps engineers and designers understand how much noise a device introduces relative to its signal.
Omni-Path is a high-performance network interconnect technology designed primarily for high-performance computing (HPC) environments. Developed by Intel, Omni-Path aims to provide enhanced scalability, efficiency, and low-latency communication compared to traditional network technologies such as InfiniBand and Ethernet.
Population statistics is a branch of statistics that focuses on the characteristics and dynamics of human populations. It involves the collection, analysis, interpretation, and presentation of data regarding population size, distribution, structure, and changes over time. This field encompasses a wide range of topics, including but not limited to: 1. **Demographic Data**: Information about the population, including age, sex, race, ethnicity, marital status, education level, and occupation.
Mel Bay is a publishing company known for producing instructional books, sheet music, and learning materials primarily for various musical instruments. Founded by guitarist Mel Bay in 1947, the company gained popularity for its guitar method books and has since expanded to cover a wide range of instruments, including banjo, mandolin, fiddle, and more. Mel Bay's instructional materials are widely used by musicians of all levels, from beginners to advanced players, and aim to promote music education and accessibility.
As of my last update in October 2023, Lisa Lix is a brand that specializes in creating a variety of lifestyle products, particularly focusing on innovative drinkware. Their offerings often include reusable cups, bottles, and other containers designed to be environmentally friendly and stylish. The products typically emphasize sustainability, functionality, and aesthetic appeal, catering to customers who value both practicality and design in their daily lives.
Supergranulation refers to a pattern of large-scale convective flow observed on the surface of the Sun. These are essentially massive, "super" sized cells of plasma that are significantly larger than the regular convective cells known as granules, which are typically about 1,000 kilometers in size. Supergranules can range from approximately 20,000 to 30,000 kilometers across and are thought to have lifetimes of several days.
In the context of physics, particularly in theoretical and mathematical physics, a "supergroup" is a generalization of a group that incorporates both commutative (bosonic) and anti-commutative (fermionic) elements. This concept arises from the study of supersymmetry, which is a theoretical framework that suggests a symmetry between bosons and fermions.
Sven Erik Jørgensen may refer to a specific individual, but without additional context, it's difficult to determine which Sven Erik Jørgensen you are referring to, as his name could belong to multiple people across various fields, such as academia, sports, or other professions. One notable individual with this name is Sven Erik Jørgensen, a prominent scientist known for his work in ecological modeling, particularly in the field of environmental science and aquatic ecosystems.
In the context of mathematical and theoretical physics, a symmetric spectrum often refers to a situation where certain properties or quantities exhibit symmetry, leading to a balanced and uniform distribution or behavior. However, the term can have specific meanings depending on the field of study. 1. **In Mathematics (especially in Spectral Theory)**: A symmetric spectrum can refer to the eigenvalues of a symmetric operator or matrix, where spectral properties are analyzed for their symmetries.
Tate's thesis generally refers to the main argument or interpretation presented by a scholar named Tate, which could pertain to various topics depending on the field of study. If you are referring to a specific individual, work, or subject area (such as art, economics, literature, etc.
Theory of Mind (ToM) refers to the ability to attribute mental states—such as beliefs, desires, intentions, and knowledge—to oneself and to others. This cognitive capability allows individuals to understand that others may have perspectives, thoughts, and feelings that differ from their own. In humans, ToM typically develops in early childhood and is considered a fundamental aspect of social cognition.
Flexible displays refer to screen technologies that can be bent, rolled, or otherwise shaped without losing functionality. These displays are typically made with materials such as organic light-emitting diodes (OLEDs), electronic paper, or liquid crystal displays (LCDs) with flexible substrates.
The Smurfs merchandising refers to the extensive range of products and promotional items associated with the Smurfs franchise, which is based on the comic series created by Belgian artist Peyo (Pierre Culliford) in the 1950s. Over the years, The Smurfs have expanded beyond comics into animated television shows, feature films, and various forms of media.
"The Unreasonable Effectiveness of Mathematics in the Natural Sciences" is an essay written by physicist Eugene Wigner, published in 1960. In this essay, Wigner discusses the seemingly miraculous ability of mathematics to describe and predict phenomena in the natural world, suggesting that the success of mathematics in explaining and modeling physical theories is surprising and profound.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
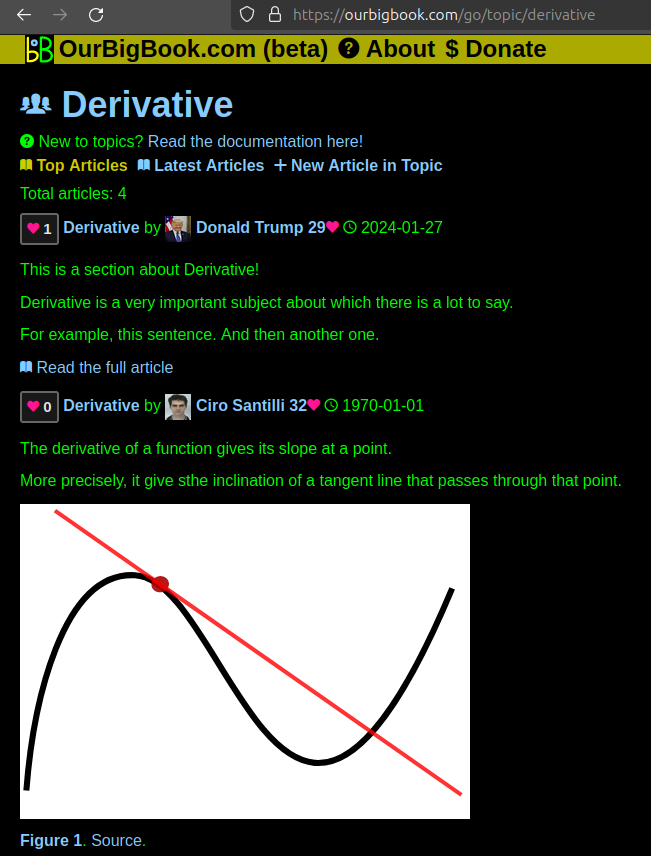
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
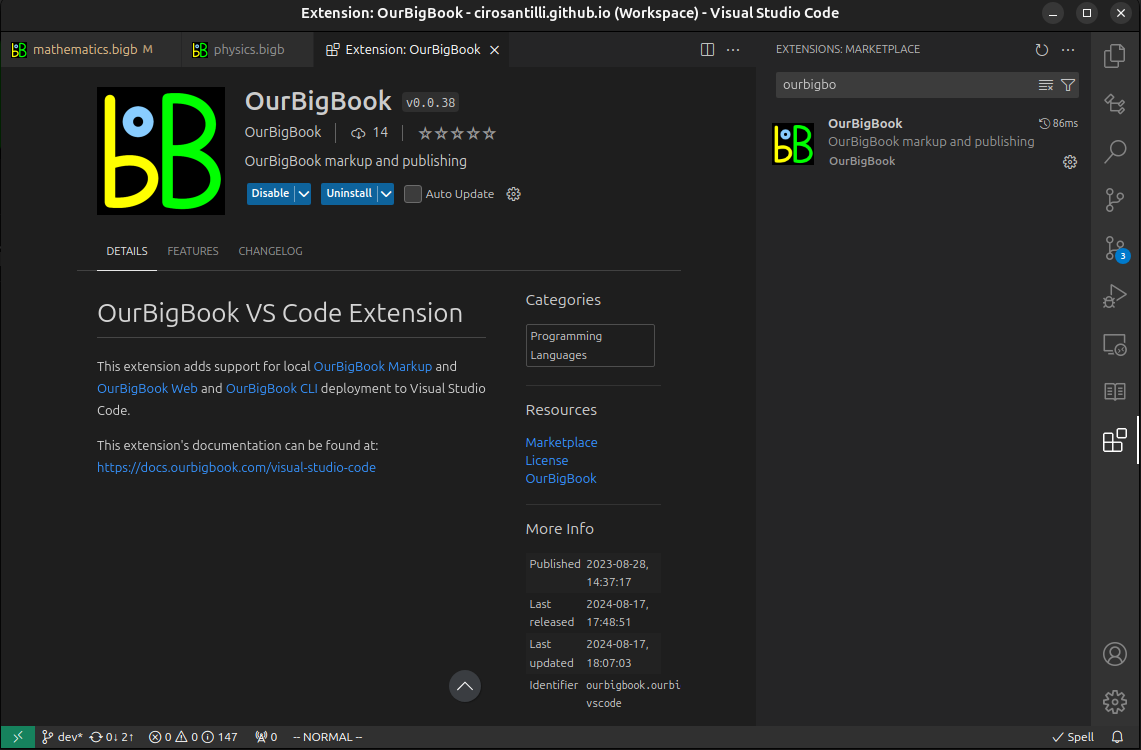
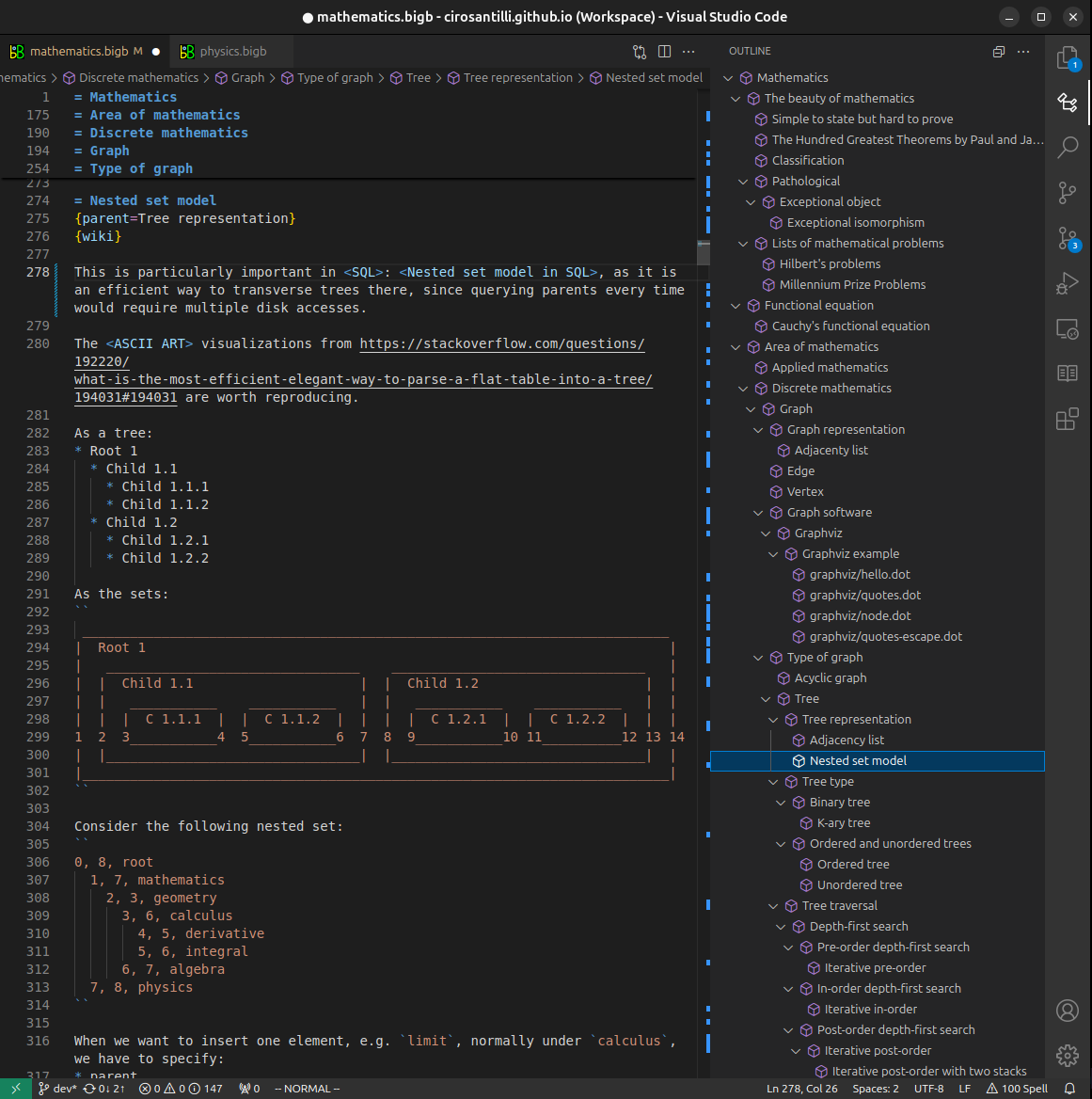
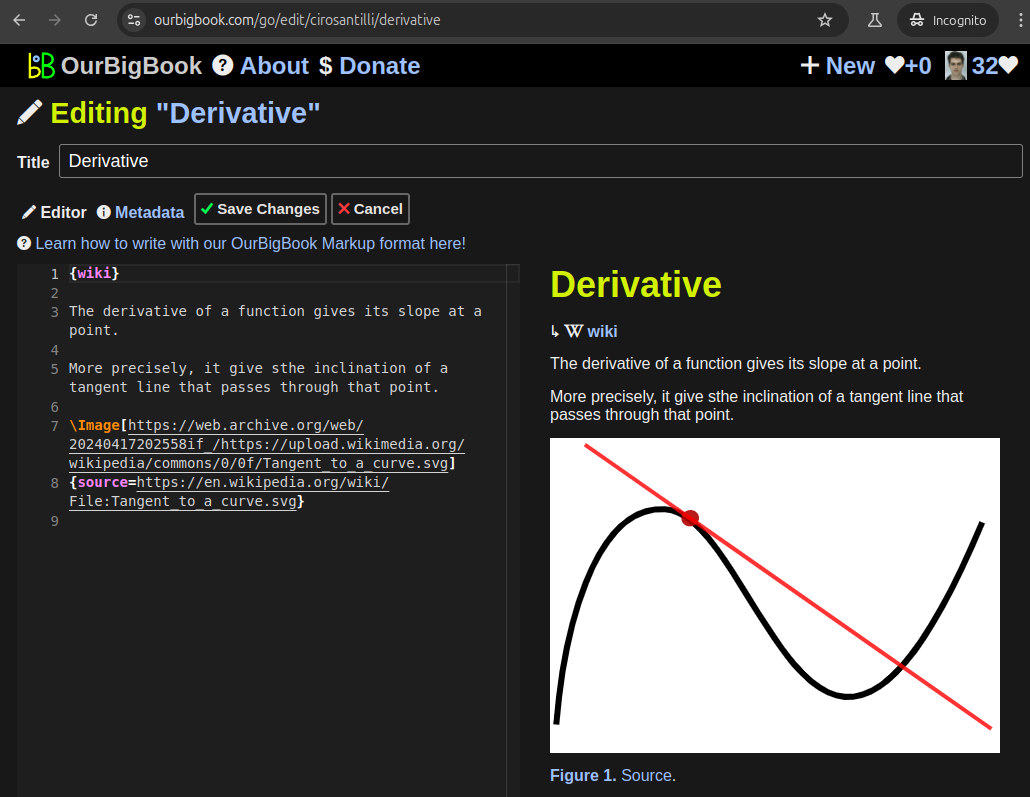
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact