Atomic and laser Physics subdepartment of the University of Oxford by  Ciro Santilli 40 Updated 2025-07-16
Ciro Santilli 40 Updated 2025-07-16
CIA 2010 covert communication websites Communication mechanism by  Ciro Santilli 40 Updated 2025-07-16
Ciro Santilli 40 Updated 2025-07-16
There are four main types of communication mechanisms found:These have short single word names with some meaning linked to their website.
- There is also one known instance where a .zip extension was used! web.archive.org/web/20131101104829*/http://plugged-into-news.net/weatherbug.zip as:
<applet codebase="/web/20101229222144oe_/http://plugged-into-news.net/" archive="/web/20101229222144oe_/http://plugged-into-news.net/weatherbug.zip"JAR is the most common comms, and one of the most distinctive, making it a great fingerprint. - JavaScript file. There are two subtypes:
- JavaScript with SHAs. Rare. Likely older. Way more fingerprintable.
- JavaScript without SHAs. They have all been obfuscated slightly different and compressed. But the file sizes are all very similar from 8kB to 10kB, and they all look similar, so visually it is very easy to detect a match with good likelyhood.
- Adobe Flash swf file. In all instances found so far, the name of the SWF matches the name of the second level domain exactly, e.g.:While this is somewhat of a fingerprint, it is worth noting that is was a relatively commonly used pattern. But it is also the rarest of the mechanisms. This is a at a dissonance with the rest of the web, which circa 2010 already had way more SWF than JAR apparently.
http://tee-shot.net/tee-shot.swfSome of the SWF websites have archives for empty/servletpages:which makes us think that it is a part of the SWF system../bailsnboots.com/20110201234509/servlet/teammate/index.html ./currentcommunique.com/20110130162713/servlet/summer/index.html ./mynepalnews.com/20110204095758/servlet/SnoopServlet/index.html ./mynepalnews.com/20110204095403/servlet/release/index.html ./www.hassannews.net/20101230175421/servlet/jordan/index.html ./zerosandonesnews.com/20110209084339/servlet/technews/index.html - CGI comms
Because the communication mechanisms are so crucial, they tend to be less varied, and serve as very good fingerprints. It is not ludicrous, e.g. identical files, but one look at a few and you will know the others.
Later on, we've also come across some stylistic hits in IP ranges with apparent slight variations of the CGI comms pattern:
Since these are so rare, it is still a bit hard to classify them for sure, but they are of great interest no doubt, as as we start to notice these patterns more tend to come if it is a thing.
Announcements and updates by self:
- 2023-06-10: initial announcements
- twitter.com/cirosantilli/status/1667532991315230720. Follow up when more domains were found: twitter.com/cirosantilli/status/1717445686214504830
- www.reddit.com/r/OSINT/comments/146185r/i_found_16_new_cia_covert_communication_websites/. Marked as SPAM 5 by mods days later. After reaching 92 votes, a very positive reply for that niche sub, and being obviously on topic. Weird. Anyways, did its job and likely kicked off hackernews.
- www.facebook.com/cirosantilli/posts/pfbid04KvRbEXghJakcD4AQz4379L5oVjPZ6vrBF1Eak3p81VnqRSXuXdvvYonCWPhGfQXl
- 2023-10-26 twitter.com/cirosantilli/status/1717445686214504830: announcement by self after finding 75 more sites
- Shared by others soo after:
- www.reddit.com/r/conspiracy/comments/14705gp/cia_2010_covert_communication_websites/ failed attempt with bad link unfortunately
- 2024-01-15: twitter.com/cirosantilli/status/1747742453778559165 Oleg Shakirov's findings
- 2024-01-23: mastodon.social/@cirosantilli/111807480628392615 ipinf.ru gives 4 hits and 4 new suspects, announced at:
- 2024-09 Aratu Week 2024 Talk by Ciro Santilli: My Best Random Projects
- 2025-03-13: 44 new domains found: Section "44 new CIA websites"
- 2025-04-14: cqcounter screenshots used to confirm many new hits: Section "60 new CIA website screenshots discovered on CQ Counter"
- 2025-05-23: Section "Backing up CIA website archives for research and posterity"
Pings by self:
- 2025-03-13:
- x.com/cirosantilli/status/1900278353065894324 pings x.com/JackRhysider Jack Rhysider, host of the Darkent Diaries podcast
- x.com/cirosantilli/status/1900828210578727276 pings x.com/JennaMC_Laugh Jenna McLaughlin and x.com/zachsdorfman Zach Dorfman, authors of the 2018 Yahoo articles
- 2025-03-31 going to find random interested people on Twitter:
- 2025-05-05:
- inteltoday.org/2021/07/31/us-national-whistleblower-day-july-30-2021-i-john-reidy-declare-cia-debacle-in-iran-china/#comment-46375 pings the author Dr. Ludwig De Braeckeleer. Besides his interest in intelligence, the dude actually also won a Breakthrough Prize in Physics, holy fuck it's mind boggling.
- x.com/cirosantilli/status/1919391488422662245 pings x.com/marisaataylor Marisa Taylor, author of the 2014, McClathy DC article
- x.com/cirosantilli/status/1919859345593880812 and x.com/cirosantilli/status/1919846838850499002 pings x.com/billmarczak Bill Marczak and x.com/thezedwards Zach Edwards, technical analysts for the Reuters article
- x.com/cirosantilli/status/1919860643408007644 pings x.com/joel_schectman Joel Schectman and x.com/bozorgmehr Bozorgmehr Sharafedin authors of the Reuters article
- x.com/cirosantilli/status/1919870831758365113 pings x.com/zachsdorfman Zach Dorfman (protected account) author of the Foreign Policy article
- x.com/cirosantilli/status/1920073080363241727 pings x.com/markmazzettinyt Mark Mazzetti, x.com/nytmike Michael S. Schmidt and x.com/mattapuzzo Matt Apuzzo authors of the 2017 New York Times article. Could not find a Twitter for the fourth author Adam Goldman.
- x.com/PeteWilliamsNBC Pete Williams author of the 2018 NCB News artricle: he's retired and not active on Twitter, not going to bother pinging
Reactions by others:
- 2023-06-19: www.reddit.com/r/numberstations/comments/14dexiu/after_numbers_stations_vanished/ (30 points) off topic on that sub, but thankfully was not deleted, interesting sub topic
- 2023-10-26: Google Analytics backlink from lms.fh-wedel.de/ path unknown. Some shitty German university: en.wikipedia.org/wiki/Fachhochschule_Wedel_University_of_Applied_Sciences LMS stands for Learning management system, apparently a Moodle instance. Maybe they have some Open educational resources, but all in German so pointless
- Second wave:
- 2023-12-01: news.ycombinator.com/item?id=38492304 (65 points). Second submission but pointing to OurBigBook.com rather than cirosantilli.com: ourbigbook.com/cirosantilli/cia-2010-covert-communication-websites We take those. Reached only 65 points as of January 2024.
- 2023-12-02: buttondown.email/grugq/archive/december-2-2023/. "grugq" is the handle of a zero day dealer whose received some scrutiny in 2012 after a Forbes protile was written about him: archive.ph/7mUG5. He comments:presumably referring to DNS Census 2013.
I don’t think anyone anticipated that databases leaked by hackers would enable OSINT researchers to conduct counterintelligence investigations that rival the state security services.
- 2024-01-12: twitter.com/jeremy_wokka/status/1745657801584656564 (40k followers, mid of thread)
- 2025-04-02: www.reddit.com/r/wikipedia/comments/1kd7rzo/comment/mqoocu7/?context=3 user Gilda1234_ mentions this project in a comment to "Between 2010 and 2012, China identified and killed at least 30 CIA informants in the country" by idlikebab
- 2025-05-26 The CIA Secretly Ran a Star Wars Fan Site by Joseph Cox from 404 media, an upstart publication covering edgy digital subjects. This was likely a result of Ciro publicly pinging x.com/zachsdorfman Zach Edwards, one of the analysts for the Reuters article, at x.com/cirosantilli/status/1919846838850499002, as he is cited in the article as having done a technical review. This had a massive knowdown effect and several other media picked the story up. Ciro announcing it at:Forum threads spawned from it:Other media that picked it up:
- Reddit
- www.reddit.com/r/StarWars/comments/1kvtzwm/the_cia_secretly_ran_a_star_wars_fan_site/
- www.reddit.com/r/StarWarsEU/comments/1kvu5g8/the_cia_secretly_ran_a_star_wars_fan_site/
- www.reddit.com/r/nottheonion/comments/1kxtpw4/the_cia_secretly_ran_a_star_wars_fan_site/
- www.reddit.com/r/nottheonion/comments/1kxtpw4/comment/muu531n/A source: www.thesun.co.uk/sport/33904606/putin-spies-cristiano-ronaldo-youtube-videos-messages/
They aren’t the only ones who do stuff like this, Russian agents were using Ronaldo highlight vids on YouTube to communicate 😭
- www.reddit.com/r/nottheonion/comments/1kxtpw4/comment/muu531n/
- www.reddit.com/r/Games/comments/1kye0pj/the_cia_operated_a_network_of_gaming_sites_and/
- www.reddit.com/r/technology/comments/1kvvx48/the_cia_secretly_ran_a_star_wars_fan_site_the/
- www.reddit.com/r/realtech/comments/1kvwigb/the_cia_secretly_ran_a_star_wars_fan_site_the/
- www.reddit.com/r/conspiracy/comments/1kw991t/the_cia_secretly_ran_a_star_wars_fan_site/
- www.reddit.com/r/LowStakesConspiracies/comments/1kwbf2y/the_cia_run_a_star_wars_fansite/
- www.reddit.com/r/andor/comments/1kw587c/very_andor_the_cia_secretly_ran_a_star_wars_fan/
- www.reddit.com/r/BrasildoB/comments/1kw6qah/um_brasileiro_acaba_de_publicar_detalhes_sobre_os/
- www.reddit.com/r/mexico/comments/1kwhuqg/la_incre%C3%ADble_web_de_star_wars_que_us%C3%B3_la_cia_para/
- www.reddit.com/r/KotakuInAction/comments/1kwoch5/the_cia_ran_a_star_wars_fan_site_to_secretly/
- www.reddit.com/r/starwarscanon/comments/1kwoxoq/til_the_cia_secretly_ran_a_star_wars_fan_site/
- www.reddit.com/r/TrueAnon/comments/1kwhxmj/cia_ran_star_wars_fan_site/
- www.reddit.com/r/MauLer/comments/1kxqim9/figures/
- www.reddit.com/r/Intelligence/comments/1kwybso/cia_uses_star_wars_website_to_communicate_with/
- www.reddit.com/r/StarWarsCirclejerk/comments/1kwzeto/in_my_mind_all_last_jedi_haters_are_feds/
- www.reddit.com/r/memes/comments/1kw5x9n/the_cia_really_gets_creative_sometimes/
- www.reddit.com/r/BurnNotice/comments/1kw4b5k/the_cia_secretly_ran_a_star_wars_fan_site_for/
- www.reddit.com/r/BurnNotice/comments/1kw4b5k/comment/muemkhk/ brings up
europeangoldfinch.netfirst described in Season 2 of Prison Break in 2007.Europeangoldfinch.net was a website used by Michael Scofield that allowed the Fox River Eight to communicate with each other on its online message board
- www.reddit.com/r/BurnNotice/comments/1kw4b5k/comment/muemkhk/ brings up
- Decent tweets:
- x.com/CultureCrave/status/1927119278047727908 600k followers
- x.com/val_reloaded/status/1927349417306161165 Argentinian Twitcher 400k followers
- news.ycombinator.com/item?id=44098274 failed unfortunately
- knockout.chat/thread/72492/1
- fanlore.org/wiki/2009-2013_CIA_communications_websites
- YouTube
Video 1. . Source. 2025-05-27. 180k subs. This one focuses on talking about the games and uses this article as the mainreference. He makes that nice note that the game Star Wars Battlefront II reached all time highs in the days following the CIA releasejThe articles apparenty coincided with the reelase of Star Wars Battlefront III alpha to lukewarm reception. Video 2. Star Wars Fan Sites Are Run by THE FEDS?! by Clownfish TV. Source. 2025-05-27. 600k subs. Video didn't take off however. The channel seems to be semi dead. But it is cool to see an American with YouTube-worth eloquence going over it.Video 3. . Source. 2025-05-28. 2M subs. He's basically reading the techspot article: www.techspot.com/news/108062-cia-used-star-wars-fan-site-secretly-communicate.html. Video 5. . Source. Seytonic had previously covered Reuters article at this other video:Video 8. . Source. 2025-06-13. 12k subs. This video draws on some research from this article, citing it on the source list: docs.google.com/document/d/1k7-YoOMRTL8qKE_FoRnyvR1QDa0JTBJo_a-SRwVMEu4/edit?tab=t.0 and using some of the screenshots.This video has some good mentions of the details of Jerry Chun Shing Lee's story which Ciro Santilli was not aware of. - other voice media:
- Meneame, a Spanish Reddit: 2025-05-27 www.meneame.net/m/tecnolog%C3%ADa/increible-web-star-wars-uso-cia-espiar-espana-mexico-otros/standard
Starting on that same day someone made starwarsweb.net redirect to cia.gov at 2025-05-26T13:28:02Z: www.whois.com/whois/starwarsweb.net- "mainstream":
- www.dailymail.co.uk/news/article-14752155/CIA-fake-websites-Star-Wars-communicate-spies.html Also announcing that:
* mastodon.social/@cirosantilli/114580297330915997
* x.com/cirosantilli/status/1927373757829583344
* www.linkedin.com/posts/cirosantilli_the-cia-secretly-ran-a-star-wars-fan-site-activity-7333140418504646656-eRzq/
* www.facebook.com/cirosantilli/posts/pfbid026kssQcXm7TwAHDJ4BQ73RKFCmJRLJsT1dfRpEmZ5GZdmsp8DukaqrbefFuGDqNZvl - www.themirror.com/news/us-news/cia-uses-star-wars-website-1174874
- www.dailymail.co.uk/news/article-14752155/CIA-fake-websites-Star-Wars-communicate-spies.html Also announcing that:
- "mainstream" non-English:
- www.france24.com/fr/%C3%A9co-tech/20250528-star-wars-bourse-ou-football-les-etranges-sites-pour-les-informateurs-de-la-cia (French)
- francais.rt.com/international/121266-espionnage-cia-utilisait-sites-fans-star-wars-pour-communiquer-secretement-avec-ses-agents-etrangers RT in French, God
- www.derstandard.at/consent/tcf/story/3000000271640/die-cia-hat-heimlich-eine-star-wars-fanseite-betrieben Der Standard (Austria)
- tw.news.yahoo.com/玩家可能都用過?原來「美國中情局」cia曾營運過遊戲媒體網站掩護行動-070742215.html (Yahoo Taiwan)
- www.abc.es/internacional/cia-empleo-sitios-web-inofensivos-paginas-star-20250528171402-nt.html (Spain)
- "non-mainstream":
- www.dexerto.com/entertainment/the-cia-secretly-used-a-star-wars-fan-site-to-talk-to-spies-report-3199318/ and x.com/Dexerto/status/1927000891963363406 large-ish publication
- www.thegamer.com/star-wars-fan-website-cia-usa-government-spies-controlled/
- www.techspot.com/news/108062-cia-used-star-wars-fan-site-secretly-communicate.html This was one of the biggest hits on Google Analytics actually.
- gigazine.net/news/20250527-starwars-fan-sites-made-by-cia/ Japanese
- Wired:
- www.pcgamer.com/gaming-industry/the-cia-operated-a-network-of-gaming-sites-and-even-a-star-wars-fanpage-that-were-part-of-one-of-its-worst-ever-intelligence-catastrophes/
- www.msn.com/en-us/news/technology/the-cia-secretly-ran-a-star-wars-fan-site-to-communicate-with-spies/ar-AA1FASFY
- www.gamespot.com/articles/the-cia-once-ran-a-star-wars-fan-site-as-part-of-a-global-intelligence-effort/1100-6532045/
- www.darkhorizons.com/how-u-s-spies-used-a-star-wars-fan-page/
- gigazine.net/news/20250527-starwars-fan-sites-made-by-cia/
- Reddit
- 2025-08-01 saw another mini-trend due to The CIA Built Hundreds of Covert Websitesby Alan Macleod: www.mintpressnews.com/cia-secret-network-885-fake-websites/290325/
This then spawned some sindicated posts:and forum threads:- www.sott.net/article/500997-The-CIA-built-hundreds-of-covert-websites-Heres-what-they-were-hiding
- scheerpost.com/2025/08/02/the-cia-built-hundreds-of-covert-websites-heres-what-they-were-hiding/
- 2025-08-02 alethonews.com/2025/08/02/the-cia-built-hundreds-of-covert-websites-heres-what-they-were-hiding/ (Greek)
- 2025-08-02 popularresistance.org/the-cia-built-hundreds-of-covert-websites/
- 2025-08-04 cz24.news/alan-macleod-cia-vytvorila-stovky-tajnych-webu-globalni-spionazni-terminaly-co-vlastne-skryvaly/ (Czech)
Notable reactions to the websites themselves:
- 2022-09-29 www.reddit.com/r/soccer/comments/xrgua4/the_cia_used_a_message_board_on_a_fake_soccer/ "The CIA used a message board on a fake soccer website called "Iraniangoals.com" to communicate with Iranian spies, dozens of whom were arrested after the website was discovered." by user Carlos-Dangerzone
Yet, all breakthroughs, comes from them, because the people who are crazy enough to believe they can change the world are the ones who actually do ;-)
Ciro Santilli intends to move his beauty list here little by little: github.com/cirosantilli/mathematics/blob/master/beauty.md
The most beautiful things in mathematics are results that are:
- simple to state but hard to prove:
- Fermat's Last Theorem
- number of unknown rationality, e.g. is rational?
- transcendental number conjectures, e.g. is transcendental?
- basically any conjecture involving prime numbers:
- many combinatorial game questions, e.g.:
- surprising results: we had intuitive reasons to believe something as possible or not, but a theorem shatters that conviction and brings us on our knees, sometimes via pathological counter-examples. General surprise themes include:Lists:
- classification of potentially infinite sets like: compact manifolds, etc.
- problems that are more complicated in low dimensions than high like:
- generalized Poincaré conjectures. It is also fun to see how in many cases complexity peaks out at 4 dimensions.
- classification of regular polytopes
- unpredictable magic constants:
- why is the lowest dimension for an exotic sphere 7?
- why is 4 the largest degree of an equation with explicit solution? Abel-Ruffini theorem
- undecidable problems, especially simple to state ones:
- mortal matrix problem
- sharp frontiers between solvable and unsolvable are also cool:
- attempts at determining specific values of the Busy beaver function for Turing machines with a given number of states and symbols
- related to Diophantine equations:
- applications: make life easier and/or modeling some phenomena well, e.g. in physics. See also: explain how to make money with the lesson
Good lists of such problems Lists of mathematical problems.
Whenever Ciro Santilli learns a bit of mathematics, he always wonders to himself:Unfortunately, due to how man books are written, it is not really possible to reach insight without first doing a bit of memorization. The better the book, the more insight is spread out, and less you have to learn before reaching each insight.
Am I achieving insight, or am I just memorizing definitions?
Here is a more understandable description of the semi-satire that follows: math.stackexchange.com/questions/53969/what-does-formal-mean/3297537#3297537
You start with a very small list of:
- certain arbitrarily chosen initial strings, which mathematicians call "axioms"
- rules of how to obtain new strings from old strings, called "rules of inference" Every transformation rule is very simple, and can be verified by a computer.
Using those rules, you choose a target string that you want to reach, and then try to reach it. Before the target string is reached, mathematicians call it a "conjecture".
Since every step of the proof is very simple and can be verified by a computer automatically, the entire proof can also be automatically verified by a computer very easily.
Finding proofs however is undoubtedly an uncomputable problem.
Most mathematicians can't code or deal with the real world in general however, so they haven't created the obviously necessary: website front-end for a mathematical formal proof system.
The fact that Mathematics happens to be the best way to describe physics and that humans can use physical intuition heuristics to reach the NP-hard proofs of mathematics is one of the great miracles of the universe.
Once we have mathematics formally modelled, one of the coolest results is Gödel's incompleteness theorems, which states that for any reasonable proof system, there are necessarily theorems that cannot be proven neither true nor false starting from any given set of axioms: those theorems are independent from those axioms. Therefore, there are three possible outcomes for any hypothesis: true, false or independent!
Some famous theorems have even been proven to be independent of some famous axioms. One of the most notable is that the Continuum Hypothesis is independent from Zermelo-Fraenkel set theory! Such independence proofs rely on modelling the proof system inside another proof system, and forcing is one of the main techniques used for this.
The landscape of modern Mathematics comic by Abstruse Goose
. Source. This comic shows that Mathematics is one of the most diversified areas of useless human knowledge.This thing is sexy.
This one strikes the right balance between restriction and permissions. NC and ND are simply too restrictive.
TODO where does the SA boundary end? E.g.:
- software: opensource.stackexchange.com/questions/173/what-do-i-need-to-share-if-i-include-cc-by-sa-artwork-in-my-software/11323#11323
- video game:
- website: graphicdesign.stackexchange.com/questions/68805/using-cc-by-sa-3-0-images-in-website-does-share-alike-affect-my-websites-lice/145124#145124
- book: academia.stackexchange.com/questions/48375/using-images-with-cc-by-sa-license-in-slides-or-a-thesis
- music in a podcast: opensource.stackexchange.com/questions/7022/using-cc-by-sa-music-in-a-podcast
White papers are a form of advertisement. They are not peer reviewed papers and are generally not reproducible; their value lies entirely in trust of their publisher rather than being able to verify their claims.
If a product of a big company has a catchy name it came from an acquisition by  Ciro Santilli 40 Updated 2025-07-16
Ciro Santilli 40 Updated 2025-07-16
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
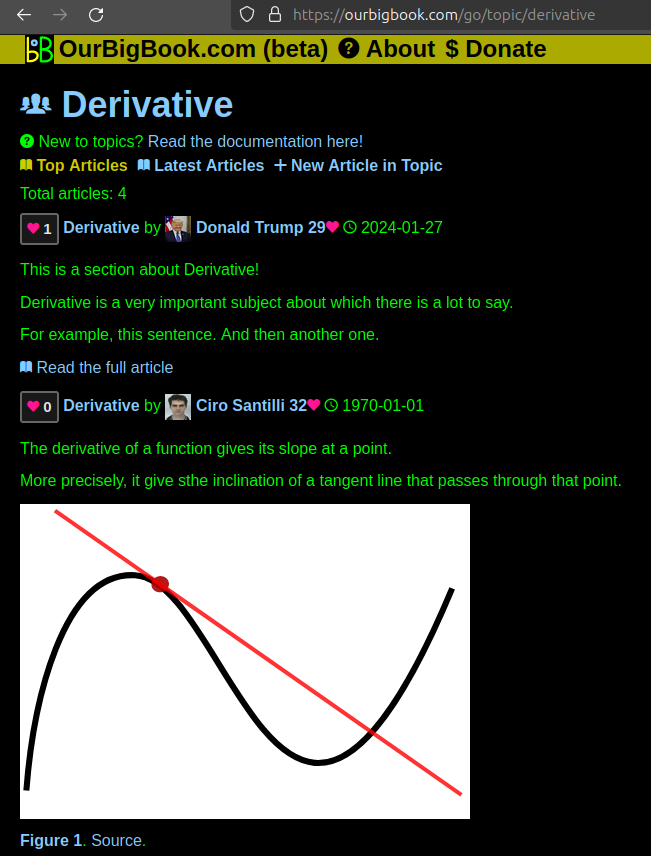
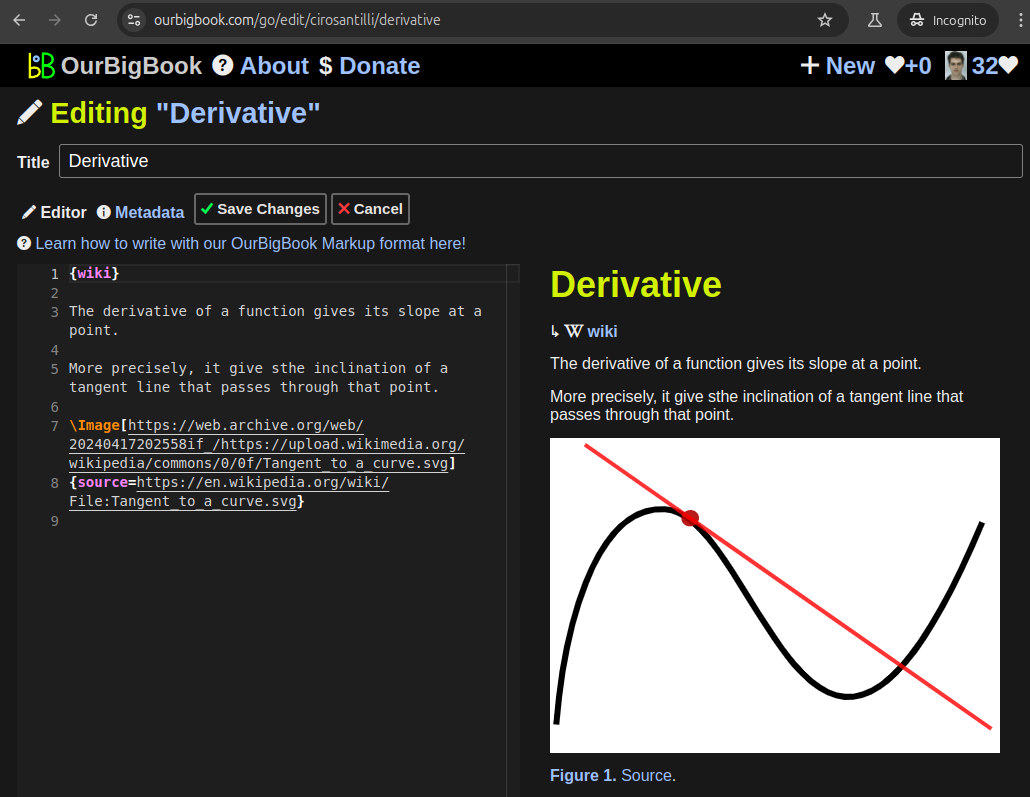
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
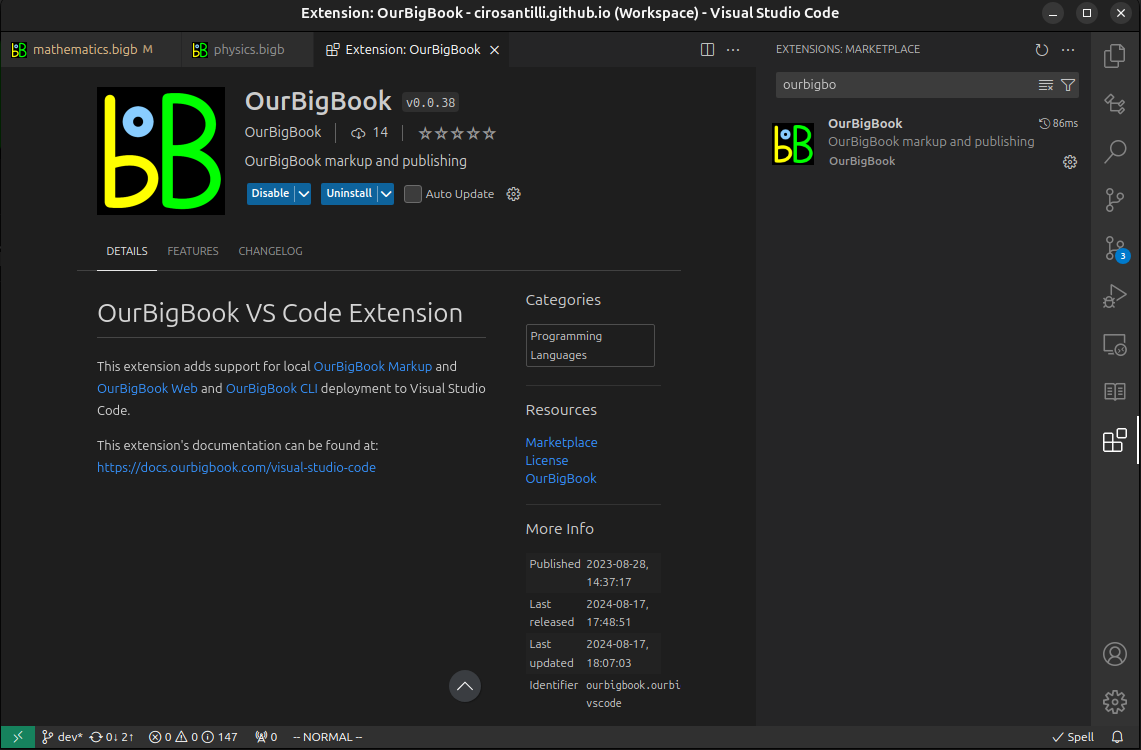
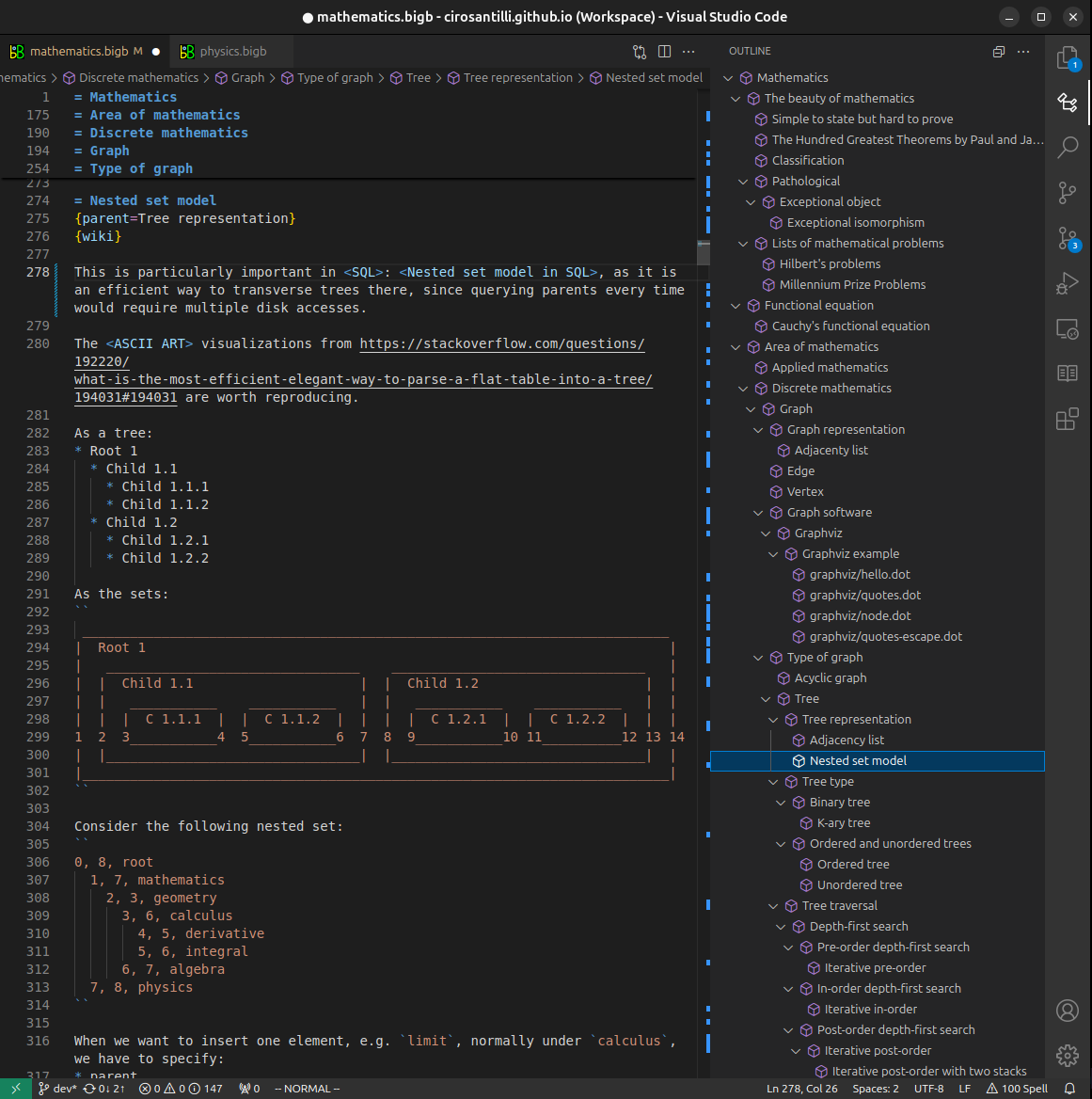
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact