The World Register of Marine Species (WoRMS) is an authoritative and comprehensive database that aims to provide valid scientific names and classification information for marine organisms. It was established to facilitate the study of marine biodiversity by offering a reliable resource for researchers, policymakers, and educators interested in marine species. Key features of WoRMS include: 1. **Taxonomic Information**: WoRMS provides taxonomic details for marine species, including synonyms, geographic distribution, and ecological information.
Zoogeography is a branch of biogeography that deals with the study of the geographical distribution of animal species and populations across the planet. It involves understanding how historical, ecological, and environmental factors influence the presence and diversity of animal life in different regions. Key areas of focus in zoogeography include: 1. **Species Distribution**: Investigating where different animal species are found and how their distributions are influenced by factors such as climate, geography, and habitat availability.
Eratosthenes Seamount is an underwater mountain located in the eastern Mediterranean Sea, off the coast of Cyprus. It is named after the ancient Greek mathematician and astronomer Eratosthenes, who is known for calculating the Earth's circumference. The seamount is part of the Eratosthenes Plateau, which is a large geological structure characterized by its complex geological history and unique bathymetry.
In oceanography, a "front" refers to a boundary or transition zone between two different water masses that have distinct physical properties, such as temperature, salinity, or density. These differences can lead to variations in water characteristics and can significantly influence marine ecosystems, weather patterns, and ocean circulation. There are several types of fronts in oceanography: 1. **Temperature Fronts**: These occur where there is a sharp change in water temperature, often associated with currents or upwelling zones.
Geophysical fluid dynamics (GFD) is a branch of fluid dynamics that focuses on the behavior of fluids in the Earth's atmosphere and oceans, as well as in other planetary environments. It combines principles from fluid mechanics, geophysics, and applied mathematics to study the motion of large-scale fluid systems influenced by the Earth's rotation, gravity, and other geophysical forces.
Jason-1 is a satellite mission that was part of a series of oceanographic satellites designed to monitor the state of the world's oceans. Launched on December 7, 2001, Jason-1 was a collaboration between NASA and the French space agency CNES (Centre National d'Études Spatiales). The primary mission of Jason-1 was to measure sea surface height, which is crucial for understanding ocean circulation, climate change, and sea level rise.
The North Atlantic Oscillation (NAO) is a weather phenomenon that is defined by fluctuations in atmospheric pressure differences between the Icelandic Low and the Azores High in the North Atlantic region. This oscillation significantly influences weather patterns in Europe and North America, affecting precipitation, temperature, and storm tracks. The NAO has two phases: 1. **Positive NAO Phase**: During this phase, the pressure difference between the Icelandic Low and the Azores High is greater than normal.
An oceanic plateau is a large, flat, elevated region of the ocean floor that rises significantly above the surrounding seafloor. These geological features are typically formed by volcanic activity and are composed primarily of basaltic lava. Oceanic plateaus are generally classified as being large in area, and they often have a relatively shallow slope.
The "Rossby whistle" refers to a phenomenon related to the dynamics of fluid motion in geophysical contexts, particularly in atmosphere and oceanography. It is associated with Rossby waves, which are large-scale waves in a rotating fluid, such as the Earth's atmosphere and oceans, caused by the Coriolis effect due to the planet's rotation.
The Tasman Outflow refers to a significant oceanic current that flows out of the Tasman Sea, which is located between Australia and New Zealand. This current plays an important role in the regional oceanography of the southwestern Pacific Ocean. Key characteristics of the Tasman Outflow include: 1. **Source of Water**: It originates from the warm, tropical waters of the Coral Sea and is influenced by the East Australian Current (EAC), which flows south along the eastern coast of Australia.
In psychophysics, "roughness" refers to a quality of texture perception that pertains to how coarse or fine a surface feels to the touch. This can involve both tactile sensations and auditory perceptions, particularly in music and sound quality. 1. **Tactile Roughness**: This aspect relates to how the skin perceives surface textures. It is influenced by factors like the size, spacing, and geometry of the surface features.
An interceptor ditch is a drainage feature designed to collect and redirect surface water runoff and subsurface water from a specific area to prevent issues like flooding, erosion, or waterlogging. It typically involves a trench or channel that can be lined or unlined, and is strategically placed to intercept and manage water flow before it reaches sensitive areas, such as structures, agricultural land, or ecosystems.
Subsidence is the gradual sinking or settling of the ground's surface. This phenomenon can occur for a variety of reasons, including natural processes and human activities. Some common causes of subsidence include: 1. **Soil Settling**: Over time, the weight of buildings, water, and other materials can cause soil and sediment to compress and settle.
The phreatic zone, also known as the saturated zone, is the area beneath the Earth's surface where all the pore spaces in soil and rock are filled with water. In this zone, water moves through the soil and rocks and is influenced by gravity and pressure. The top boundary of the phreatic zone is called the water table, which separates it from the vadose zone (the unsaturated zone above the water table where the soil pores contain both air and water).
No. 1409 Flight RAF is a unit of the Royal Air Force (RAF) that was formed to provide training to pilots, specifically focusing on the development of skills necessary for operating aircraft in challenging conditions or for specific missions. As a flight within the RAF, it has had various roles over time, often associated with specific aircraft types or operational requirements. 1409 Flight has historically been involved in diverse functions, including search and rescue, reconnaissance, and pilot training.
A weather balloon is a type of high-altitude balloon specifically designed to collect data about the atmosphere. Typically made of latex or synthetic rubber, these balloons are filled with helium or hydrogen gas and can ascend to altitudes of up to 40 kilometers (about 25 miles) or more. Weather balloons are equipped with scientific instruments, such as radiosondes, which measure various atmospheric parameters, including temperature, humidity, atmospheric pressure, and wind speed and direction.
The planetary boundary layer (PBL) is a key part of the Earth's atmosphere that is directly influenced by the presence of the Earth's surface. It is the lowest portion of the atmosphere, typically extending from the surface up to about 1 to 2 kilometers (or approximately 0.6 to 1.2 miles) in altitude, although its thickness can vary depending on weather conditions, terrain, and time of day.
The term "surface layer" can refer to several contexts across different fields, including geology, oceanography, atmospheric science, and materials science. Here are a few interpretations based on these contexts: 1. **Geology and Soil Science**: In this context, the surface layer refers to the uppermost layer of soil or rock that is in contact with the environment. It is often the layer where biological activity is greatest and is critical for plant growth.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
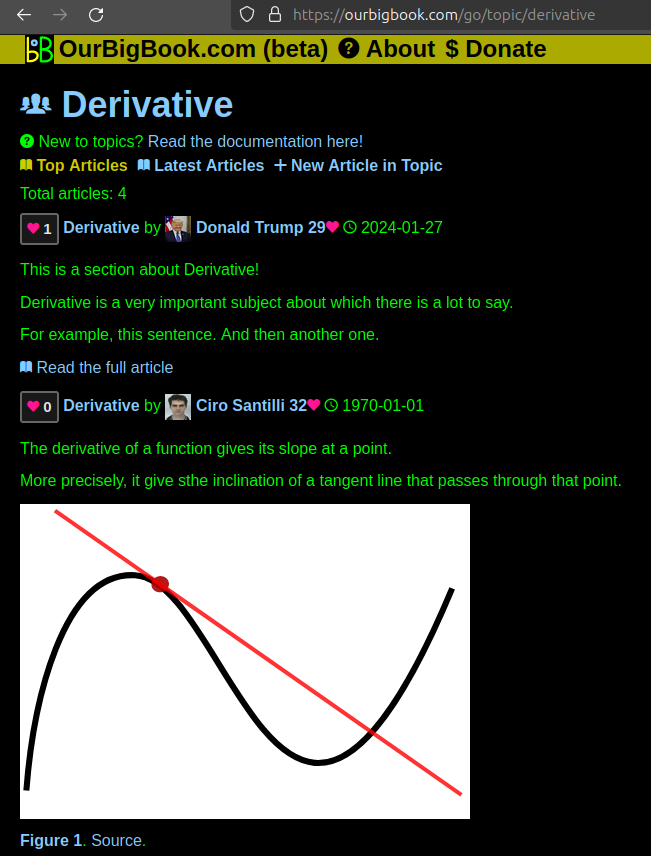
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
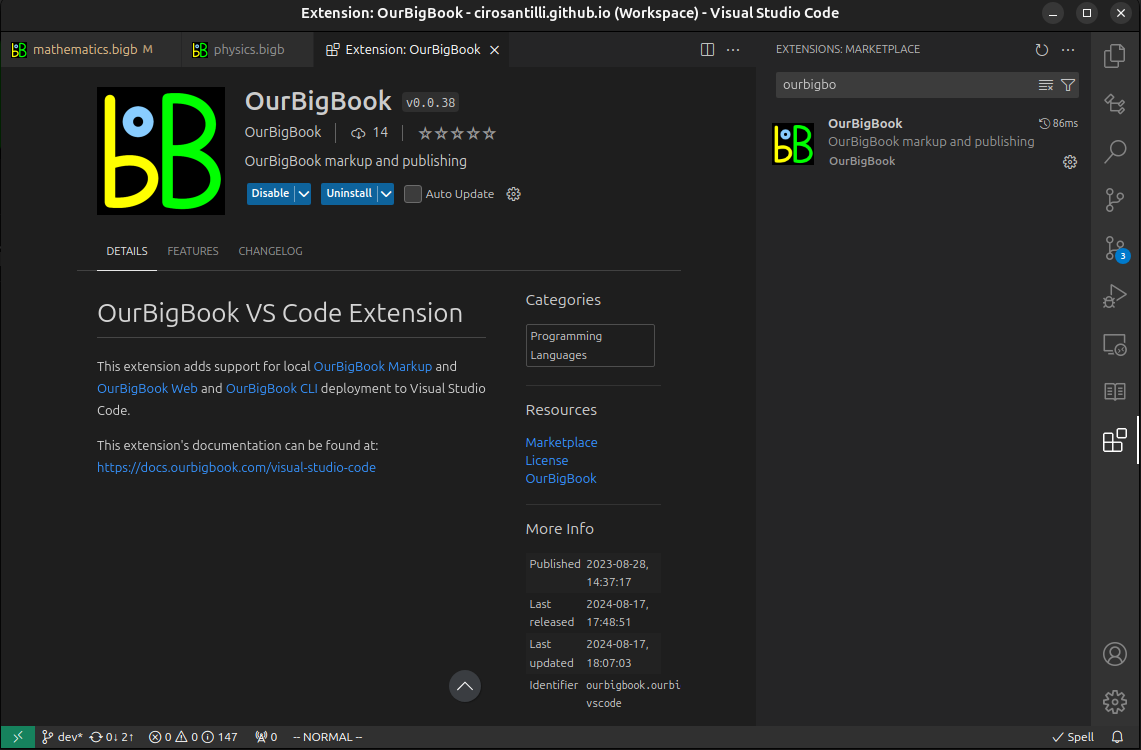
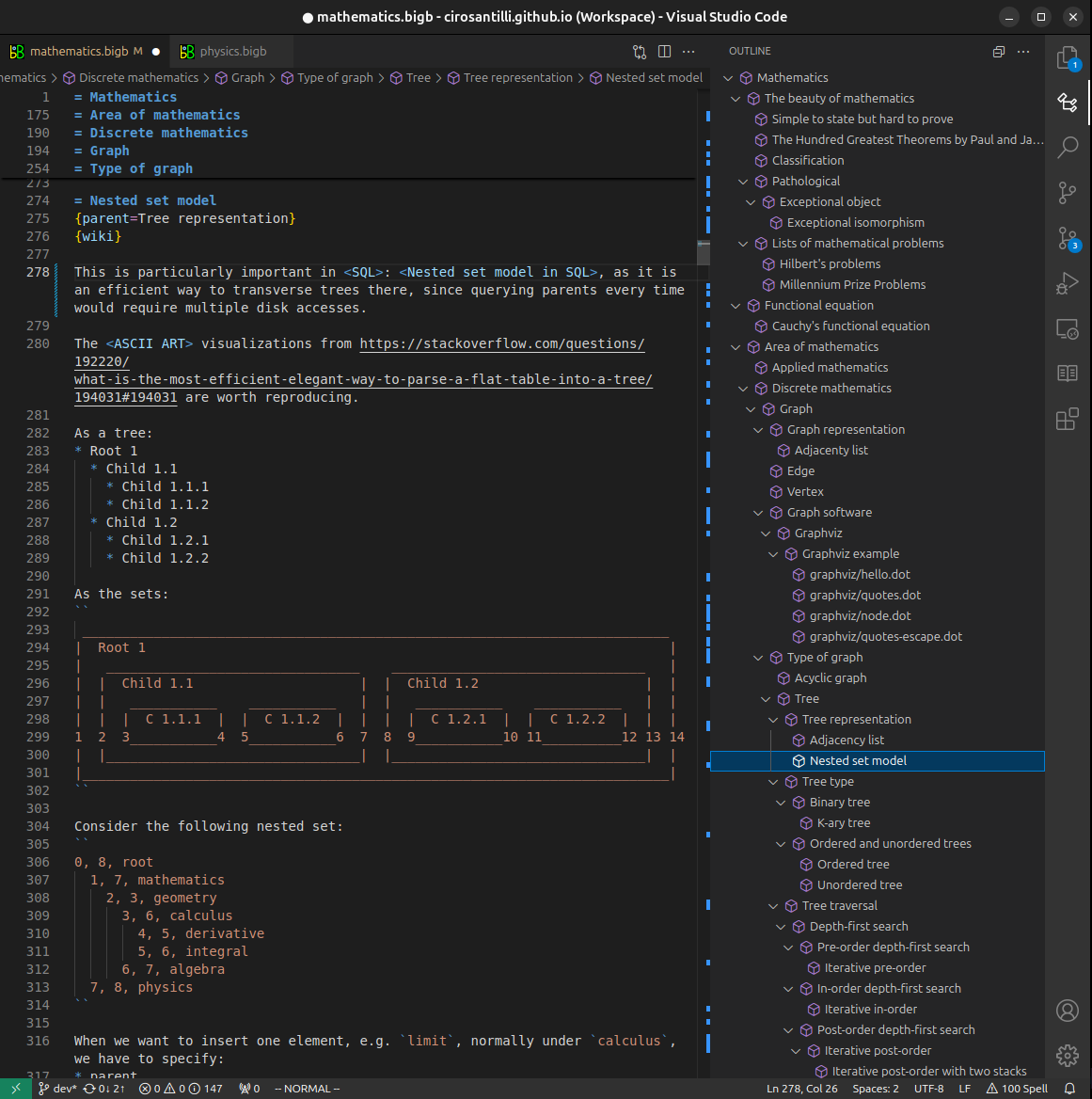
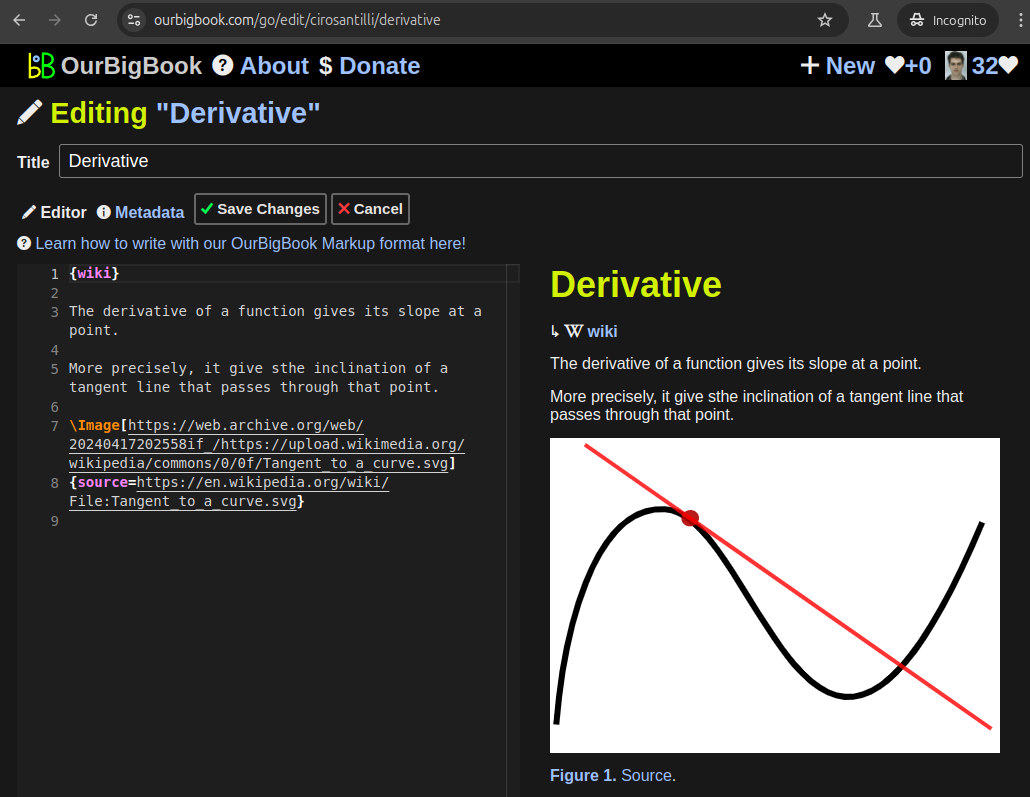
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact