As of 2021, their location is a small business park in Haywards Heath, about 15 minutes north of Brighton[ref]
Funding rounds:
- 2022:
- 67m euro contract with the German government: www.uktech.news/deep-tech/universal-quantum-german-contract-20221102 Both co-founders are German. They then immediatly announced several jobs in Hamburg: apply.workable.com/universalquantum/?lng=en#jobs so presumably linked to the Hamburg University of Technology campus of the German Aerospace Center.
- medium.com/@universalquantum/universal-quantum-wins-67m-contract-to-build-the-fully-scalable-trapped-ion-quantum-computer-16eba31b869e
- 2021: $10M (7.5M GBP) grant from the British Government: www.uktech.news/news/brighton-universal-quantum-wins-grant-20211105This grant is very secretive, very hard to find any other information about it! Most investment trackers are not listing it.The article reads:Interesting!
Universal Quantum will lead a consortium that includes Rolls-Royce, quantum developer Riverlane, and world-class researchers from Imperial College London and The University of Sussex, among others.
A but further down the article gives some more information of partners, from which some of the hardware vendors can be deduced:The consortium includes end-user Rolls-Royce supported by the Science and Technology Facilities Council (STFC) Hartree Centre, quantum software developer Riverlane, supply chain partners Edwards, TMD Technologies (now acquired by Communications & Power Industries (CPI)) and Diamond Microwave
- Edwards is presumably Edwards Vacuum, since we know that trapped ion quantum computers rely heavily on good vacuum systems. Edwards Vacuum is also located quite close to Universal Quantum as of 2022, a few minutes drive.
- TMD Technologies is a microwave technology vendor amongst other things, and we know that microwaves are used e.g. to initialize the spin states of the ions
- Diamond Microwave is another microwave stuff vendor
www.riverlane.com/news/2021/12/riverlane-joins-7-5-million-consortium-to-build-error-corrected-quantum-processor/ gives some more details on the use case provided by Rolls Royce:The work with Rolls Royce will explore how quantum computers can develop practical applications toward the development of more sustainable and efficient jet engines.This starts by applying quantum algorithms to take steps to toward a greater understanding of how liquids and gases flow, a field known as 'fluid dynamics'. Simulating such flows accurately is beyond the computational capacity of even the most powerful classical computers today.This funding was part of a larger quantum push by the UKNQTP: www.ukri.org/news/50-million-in-funding-for-uk-quantum-industrial-projects/ - 2020: $4.5M (3.5M GBP) www.crunchbase.com/organization/universal-quantum. Just out of stealth.
Co-founders:
- Sebastian Weidt. He is German, right? Yes at youtu.be/SwHaJXVYIeI?t=1078 from Video 3. "Fireside Chat with with Sebastian Weidt by Startup Grind Brighton (2022)". The company was founded by two Germans from Essex!
- Winfried Hensinger: if you saw him on the street, you'd think he plays in a punk-rock band. That West Berlin feeling.
Homepage points to foundational paper: www.science.org/doi/10.1126/sciadv.1601540
Universal Quantum emerges out of stealth by University of Sussex (2020)
Source. Explains that a more "traditional" trapped ion quantum computer would user "pairs of lasers", which would require a lot of lasers. Their approach is to try and do it by applying voltages to a microchip instead.- youtu.be/rYe9TXz35B8?t=127 shows some 3D models. It shows how piezoelectric actuators are used to align or misalign some plates, which presumably then determine conductivity
Quantum Computing webinar with Sebastian Weidt by Green Lemon Company (2020)
Source. The sound quality is to bad to stop and listen to, but it presumaby shows the coding office in the background.Fireside Chat with with Sebastian Weidt by Startup Grind Brighton (2022)
Source. Very basic target audience:- youtu.be/SwHaJXVYIeI?t=680 we are not at a point where you can buy victory. There is too much uncertainty involved across different approaches.
- youtu.be/SwHaJXVYIeI?t=949 his background
- youtu.be/SwHaJXVYIeI?t=1277 difference between venture capitalists in different countries
- youtu.be/SwHaJXVYIeI?t=1535 they are 33 people now. They've just setup their office in Haywards Heath, north of Bristol.
These people are cool.
They use optical tweezers to place individual atoms floating in midair, and then do stuff to entangle their nuclear spins.
- 2022: $15 million www.orcacomputing.com/blog/orca-computing-completes-15-million-series-a-funding-round
- 2021: $14.5 million for an Innovate UK project
CEO: Jeremy O'Brien
Raised 215M in 2020: www.bloomberg.com/news/articles/2020-04-06/quantum-computing-startup-raises-215-million-for-faster-device
Good talk by CEO before starting the company which gives insight on what they are very likely doing: Video "Jeremy O'Brien: "Quantum Technologies" by GoogleTechTalks (2014)"
PsiQuantum appears to be particularly secretive, even more than other startups in the field.
They want to reuse classical semiconductor fabrication technologies, notably they have close ties to GlobalFoundries.
Jeremy O'Brien told his peers that he had the best tech, and that he should get it all.
Some well connected peers from well known universities did not agree however, and also bid for the money, and won.
Jeremy was defeated. And pissed.
Makes for a reasonable the old man lost his horse.
www.ft.com/content/afc27836-9383-11e9-aea1-2b1d33ac3271 British quantum computing experts leave for Silicon Valley talks a little bit about them leaving, but nothing too juicy. They were called PsiQ previously apparently.More interestingly, the article mentions that this was party advised by early investor Hermann Hauser, who is known to be preoccupied about UK's ability to create companies. Of course, European Tower of Babel.
The departure of some of the UK’s leading experts in a potentially revolutionary new field of technology will raise fresh concerns over the country’s ability to develop industrial champions in the sector.
Rounds:
www.youtube.com/watch?v=v7iAqcFCTQQ shows their base technology:
- laser beam comes in
- input set via of optical ring resonators that form a squeezed state of light. Does not seem to rely on single photon production and detection experiments?
Does not appear to refer to any one specific phylogenetic level, it usually refers to either:
- Saccharomyces cerevisiae, the model eukaryote unicellular organism
- two phila of the fungus kingdom
Other good lists:
- quantumcomputingreport.com/resources/tools/ is hard to beat as usual.
- www.quantiki.org/wiki/list-qc-simulators
- JavaScript
- algassert.com/quirk demo: github.com/Strilanc/Quirk drag-and-drop, by a 2019-quantum-computing-Googler, impressive. You can create gates. State store in URL.
- github.com/stewdio/q.js/ demo: quantumjavascript.app/
Bibliography:
- www.epcc.ed.ac.uk/whats-happening/articles/energy-efficient-quantum-computing-simulations mentions two types of quantum computer simulation:
The most common approach to quantum simulations is to store the whole state in memory and to modify it with gates in a given order
However, there is a completely different approach that can sometimes eliminate this issue - tensor networks
Sample program output,
counts are randomized each time.First we take the quantum state vector immediately after the input.We understand that the first element of
input:
state:
Statevector([1.+0.j, 0.+0.j, 0.+0.j, 0.+0.j],
dims=(2, 2))
probs:
[1. 0. 0. 0.]Statevector is , and has probability of 1.0.Next we take the state after a Hadamard gate on the first qubit:We now understand that the second element of the
h:
state:
Statevector([0.70710678+0.j, 0.70710678+0.j, 0. +0.j,
0. +0.j],
dims=(2, 2))
probs:
[0.5 0.5 0. 0. ]Statevector is , and now we have a 50/50 propabability split for the first bit.Then we apply the CNOT gate:which leaves us with the final .
cx:
state:
Statevector([0.70710678+0.j, 0. +0.j, 0. +0.j,
0.70710678+0.j],
dims=(2, 2))
probs:
[0.5 0. 0. 0.5]Then we print the circuit a bit:
qc without measure:
┌───┐
q_0: ┤ H ├──■──
└───┘┌─┴─┐
q_1: ─────┤ X ├
└───┘
c: 2/══════════
qc with measure:
┌───┐ ┌─┐
q_0: ┤ H ├──■──┤M├───
└───┘┌─┴─┐└╥┘┌─┐
q_1: ─────┤ X ├─╫─┤M├
└───┘ ║ └╥┘
c: 2/═══════════╩══╩═
0 1
qasm:
OPENQASM 2.0;
include "qelib1.inc";
qreg q[2];
creg c[2];
h q[0];
cx q[0],q[1];
measure q[0] -> c[0];
measure q[1] -> c[1];This is an example of the
qiskit.circuit.library.QFT implementation of the Quantum Fourier transform function which is documented at: docs.quantum.ibm.com/api/qiskit/0.44/qiskit.circuit.library.QFTOutput:So this also serves as a more interesting example of quantum compilation, mapping the
init: [1, 0, 0, 0, 0, 0, 0, 0]
qc
┌──────────────────────────────┐┌──────┐
q_0: ┤0 ├┤0 ├
│ ││ │
q_1: ┤1 Initialize(1,0,0,0,0,0,0,0) ├┤1 QFT ├
│ ││ │
q_2: ┤2 ├┤2 ├
└──────────────────────────────┘└──────┘
transpiled qc
┌──────────────────────────────┐ ┌───┐
q_0: ┤0 ├────────────────────■────────■───────┤ H ├─X─
│ │ ┌───┐ │ │P(π/2) └───┘ │
q_1: ┤1 Initialize(1,0,0,0,0,0,0,0) ├──────■───────┤ H ├─┼────────■─────────────┼─
│ │┌───┐ │P(π/2) └───┘ │P(π/4) │
q_2: ┤2 ├┤ H ├─■─────────────■──────────────────────X─
└──────────────────────────────┘└───┘
Statevector([0.35355339+0.j, 0.35355339+0.j, 0.35355339+0.j,
0.35355339+0.j, 0.35355339+0.j, 0.35355339+0.j,
0.35355339+0.j, 0.35355339+0.j],
dims=(2, 2, 2))
init: [0.0, 0.35355339059327373, 0.5, 0.3535533905932738, 6.123233995736766e-17, -0.35355339059327373, -0.5, -0.35355339059327384]
Statevector([ 7.71600526e-17+5.22650714e-17j,
1.86749130e-16+7.07106781e-01j,
-6.10667421e-18+6.10667421e-18j,
1.13711443e-16-1.11022302e-16j,
2.16489014e-17-8.96726857e-18j,
-5.68557215e-17-1.11022302e-16j,
-6.10667421e-18-4.94044770e-17j,
-3.30200457e-16-7.07106781e-01j],
dims=(2, 2, 2))QFT gate to Qiskit Aer primitives.If we don't
transpile in this example, then running blows up with:qiskit_aer.aererror.AerError: 'unknown instruction: QFT'The second input is:and the output of that approximately:which can be defined simply as the normalized DFT of the input quantum state vector.
[0, 1j/sqrt(2), 0, 0, 0, 0, 0, 1j/sqrt(2)]From this we see that the Quantum Fourier transform is equivalent to a direct discrete Fourier transform on the quantum state vector, related: physics.stackexchange.com/questions/110073/how-to-derive-quantum-fourier-transform-from-discrete-fourier-transform-dft
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
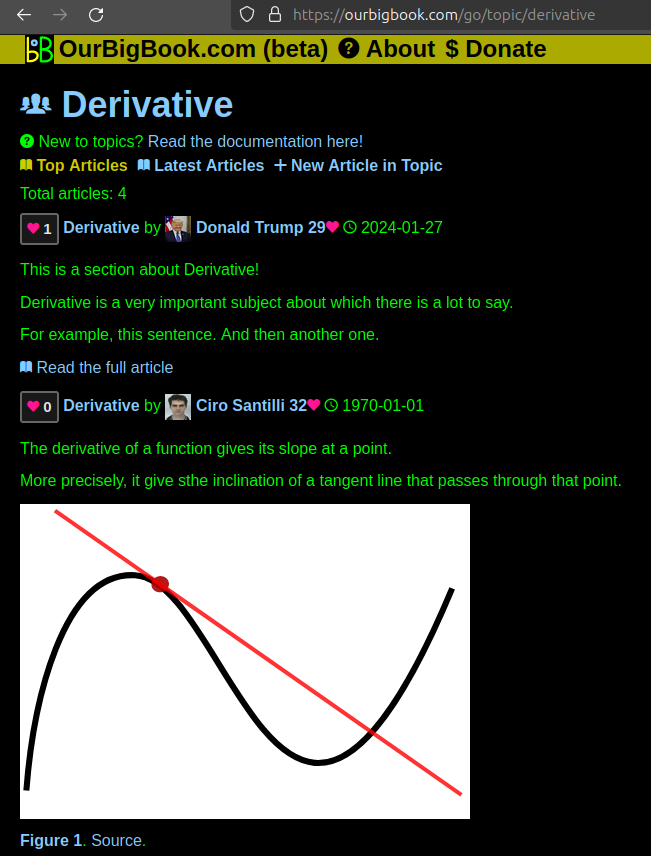
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
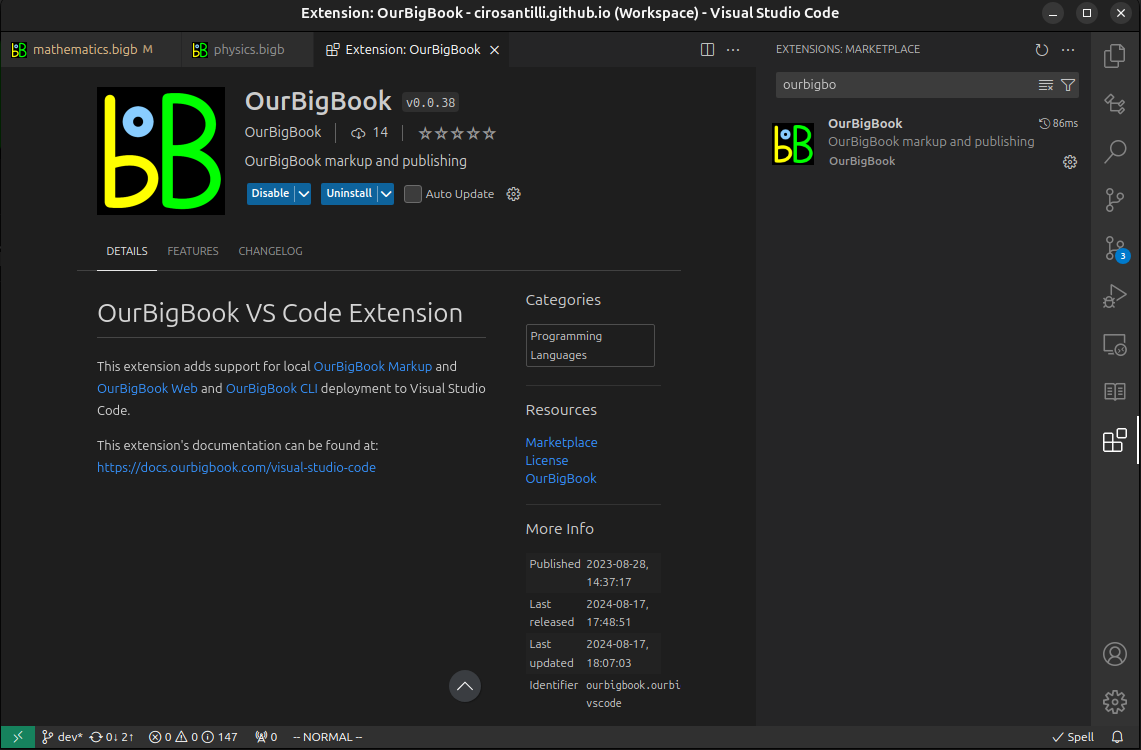
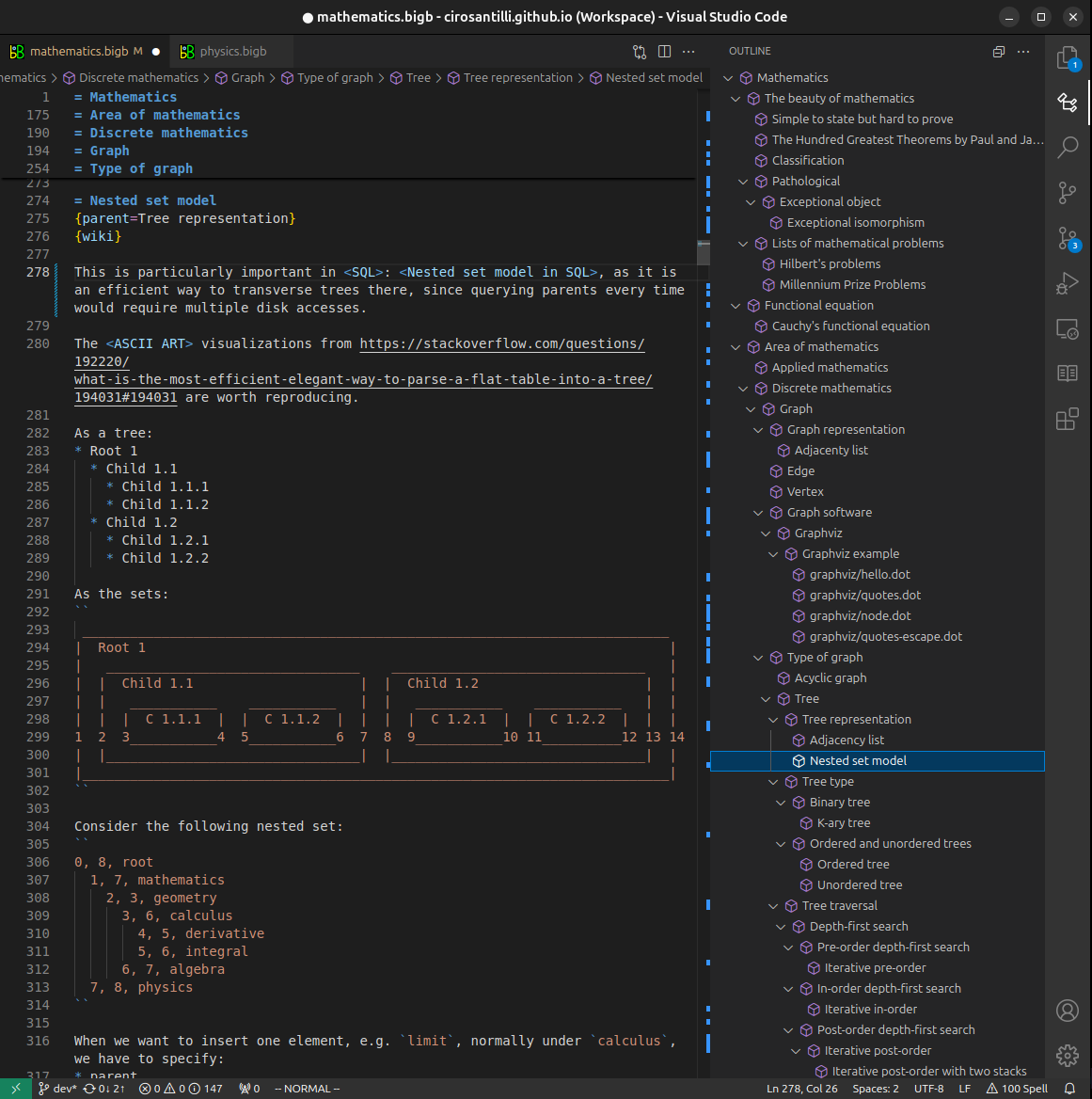
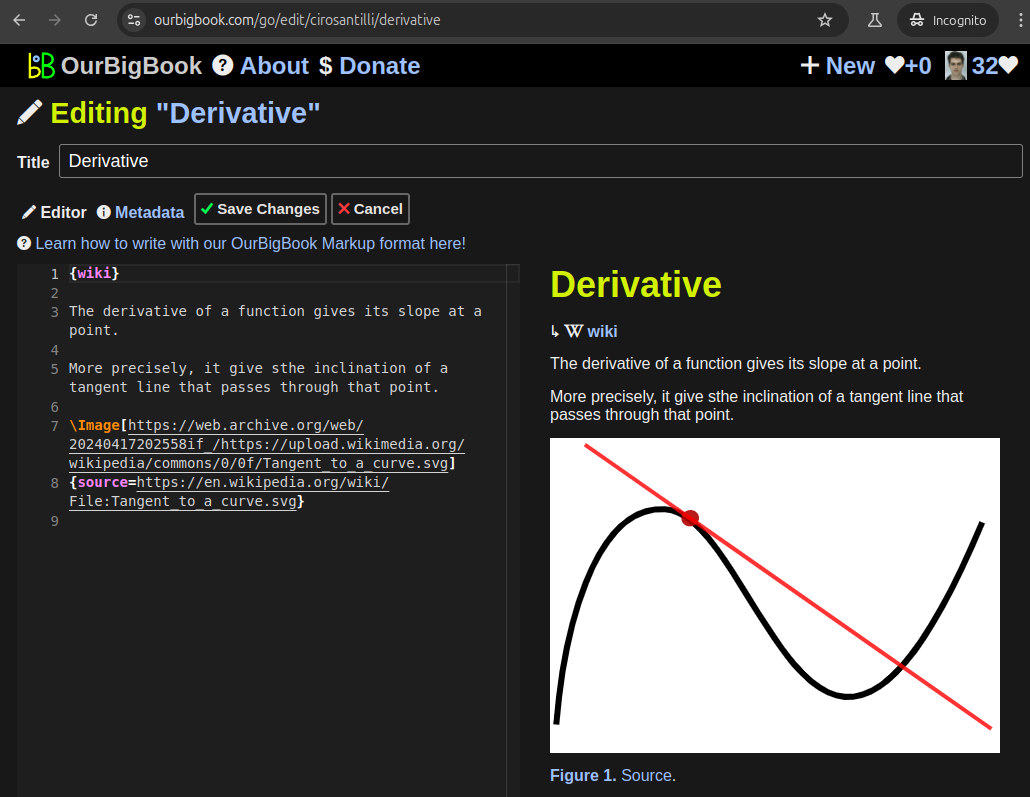
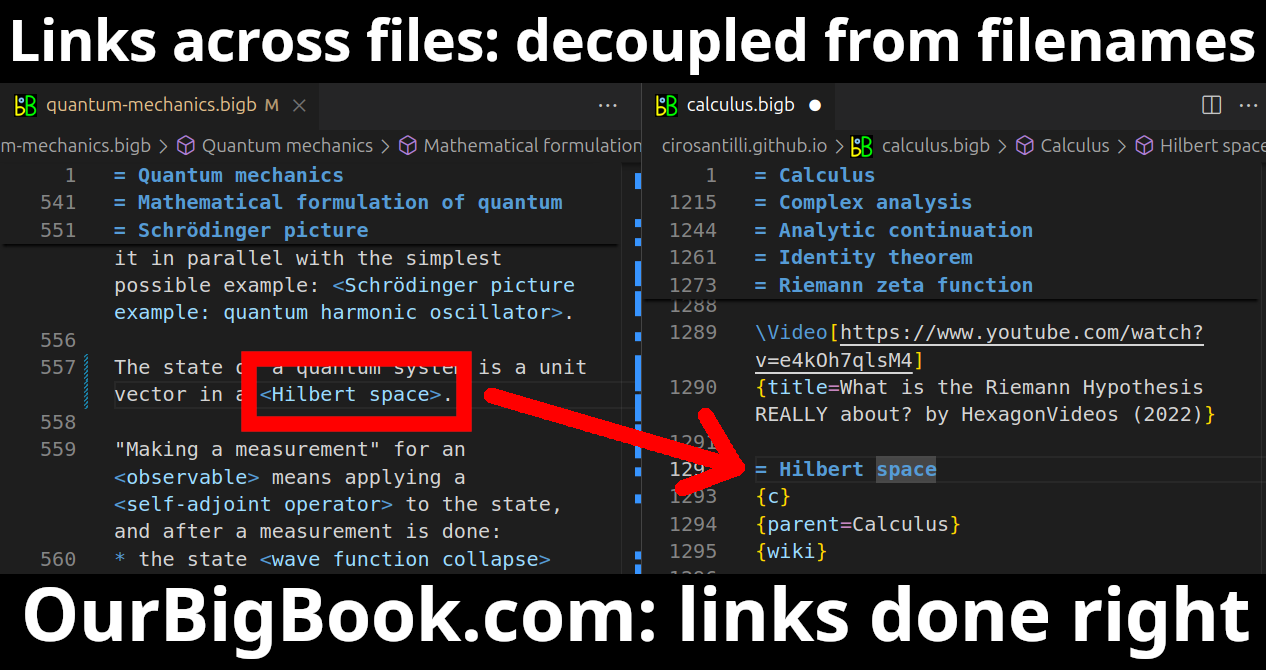
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact