As of my last knowledge update in October 2023, "PlayNET" can refer to different things depending on the context. However, there isn't a widely recognized platform or service specifically known as "PlayNET" that stands out in mainstream media or technology. It could potentially refer to: 1. **Gaming Platforms**: It might be a gaming network, community, or service related to online gaming.
Prestel, or Prestel AG, is a company that was a pioneer in the field of online services and electronic publishing. Founded in the 1970s in Germany, it became known for its interactive multimedia services that provided users with access to information and digital content via telephone lines, using specialized terminals or modems. Prestel was part of the early wave of online services that emerged before the widespread adoption of the World Wide Web.
Dow Jones News/Retrieval is a comprehensive news and information service provided by Dow Jones & Company. It allows users to access a wide variety of financial news, company information, and data from multiple sources, including newspapers, websites, and other media outlets. The service is designed primarily for financial professionals, analysts, and researchers who need timely and accurate information to make informed decisions.
The Electronic Information Exchange System (EIES) typically refers to a digital framework or platform designed to facilitate the electronic transfer and sharing of information between different entities, such as organizations, government bodies, or within specific sectors like healthcare, finance, or government services.
GameLine was a service introduced in the early 1980s that allowed users to download games and other content directly to their Atari 2600 home video game console via a telephone line. This innovation was one of the first examples of online gaming or downloading games from a remote source directly to a gaming system. The service offered a library of games that players could access by connecting their Atari console to a GameLine modem.
GreenNet is a UK-based internet service provider (ISP) that focuses on providing internet services with an emphasis on sustainability and social responsibility. Founded in 1994, GreenNet aims to offer ethical and environmentally-friendly alternatives to conventional ISPs. The company is known for serving a diverse range of clients, including non-profit organizations, social enterprises, and individuals who prioritize ethical practices.
Francisco Bezanilla may refer to a few different contexts, but it appears you may be asking about a specific individual, possibly a scientist or researcher known primarily for work in a certain field, such as biology or physiology. However, there is no widely recognized figure by that name in commonly available sources up to October 2023.
Singapore Teleview is not widely recognized as a major service or product, so it's possible that the term may refer to a specific company, service, or platform that is either lesser-known or a niche offering in Singapore. It might involve telecommunications, broadcasting, or a service related to television viewing. For the latest and most accurate information, please consult recent sources or specific local databases.
Telecom Gold was a pioneering online service that provided dial-up access to email, news, and other online services during the late 1980s and early 1990s. It was originally launched by British Telecom in 1984, primarily targeting business users but later expanded its offerings to consumers. The service allowed users to send and receive emails, access bulletin board systems, and connect with other users through various online forums.
The Source is an online platform that provides a variety of content, with a focus on news, articles, and information catering primarily to the African American community. It covers a range of topics including culture, politics, entertainment, and lifestyle. Originally launched as a print magazine in 1988, The Source has evolved over the years to establish a significant online presence, offering digital content that engages with current events and trends that resonate with its audience.
Metaculus is an online platform that focuses on forecasting and prediction markets. It allows users to make predictions about future events and outcomes across various domains, including science, technology, politics, and economics. The community of forecasters contributes their insights, and the platform aggregates these predictions to provide collective forecasts and probabilities about specific events. Metaculus operates based on a points system, where users earn points for accurate predictions, encouraging participation and engagement.
Edward H. Egelman is a prominent American biophysicist and structural biologist known for his work on the structure and function of macromolecules and molecular complexes, particularly in the context of the cytoskeleton, muscle proteins, and the interactions between proteins and nucleic acids. He has contributed significantly to the understanding of the molecular basis of various biological processes.
Ken A. Dill is a prominent biophysicist and professor known for his contributions to the field of protein folding and thermodynamics. He has worked extensively on understanding how proteins attain their functional shapes and the forces that govern these processes. Dill has made significant contributions to theoretical and computational methods in biophysics, and he has been involved in research related to protein design and molecular simulations.
Frederic M. Richards is a prominent figure in the field of biochemistry and structural biology, particularly known for his work on the structures and functions of proteins and nucleic acids. He has contributed significantly to the understanding of protein folding and stability, as well as the role of water in biological systems. Richards has also been involved in various initiatives related to scientific education and research.
André-Louis Danjon (1890–1960) was a notable French astronomer known for his contributions to astrophysics and geophysics. He is recognized for his work in various fields, including the study of the Moon, stellar photometry, and the development of photometric methods. One of his significant contributions was the establishment of the Danjon scale, which is used to classify the brightness of the Moon during lunar eclipses.
Gurtam is a technology company that specializes in software development for the telematics, logistics, and fleet management industries. Founded in 2002 and headquartered in Minsk, Belarus, Gurtam provides a range of solutions for businesses to track and manage their vehicles, assets, and workforce using GPS and IoT technologies. The company's flagship product is Wialon, a cloud-based platform that offers real-time tracking, data analytics, and reporting tools for fleet management, transportation, and logistics.
Silvia Torres-Peimbert is a prominent Mexican astrophysicist known for her contributions to the study of planetary nebulae and stellar evolution. She has held significant academic positions, including serving as a professor and researcher at the National Autonomous University of Mexico (UNAM). Torres-Peimbert is recognized for her work in promoting science education and encouraging the participation of women in the sciences.
A boiler is a closed vessel in which water or another fluid is heated. The heated fluid is then used for various purposes, such as generating steam for power generation, heating buildings, or supplying hot water for various industrial processes. **Key components and concepts related to boilers include:** 1. **Types of Boilers**: - **Fire-tube Boilers**: Hot gases from combustion pass through tubes that are surrounded by water.
Active fuel length usually refers to the portion of a nuclear reactor's fuel that is actively participating in the fission process. In a nuclear reactor, fuel rods contain nuclear fuel (typically enriched uranium or plutonium), and the length of the fuel that is effectively contributing to the chain reactions can be critical for determining the reactor's performance and efficiency. The concept is essential in the design and operation of reactors, as it affects factors like the reactor's power output, neutron economy, and overall lifecycle.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
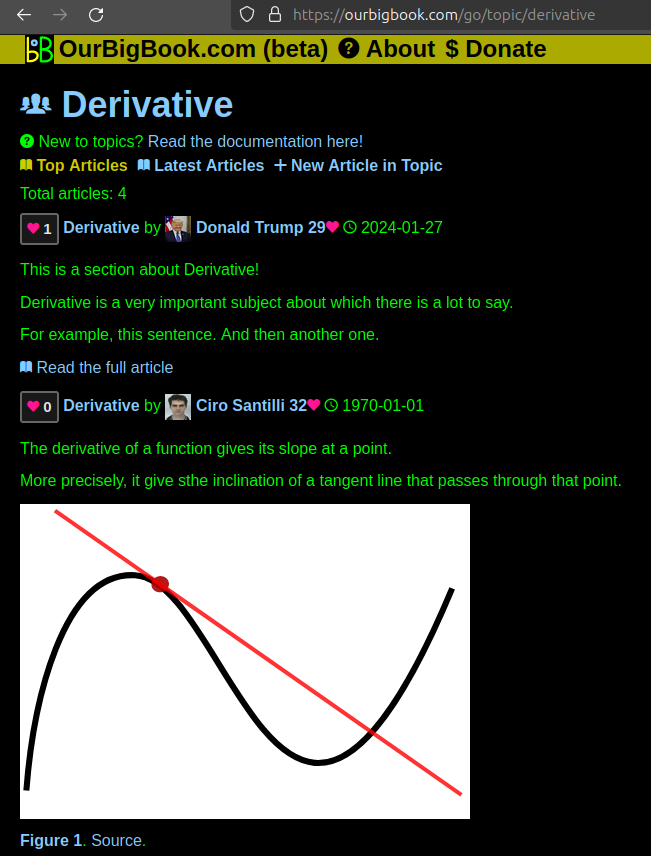
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
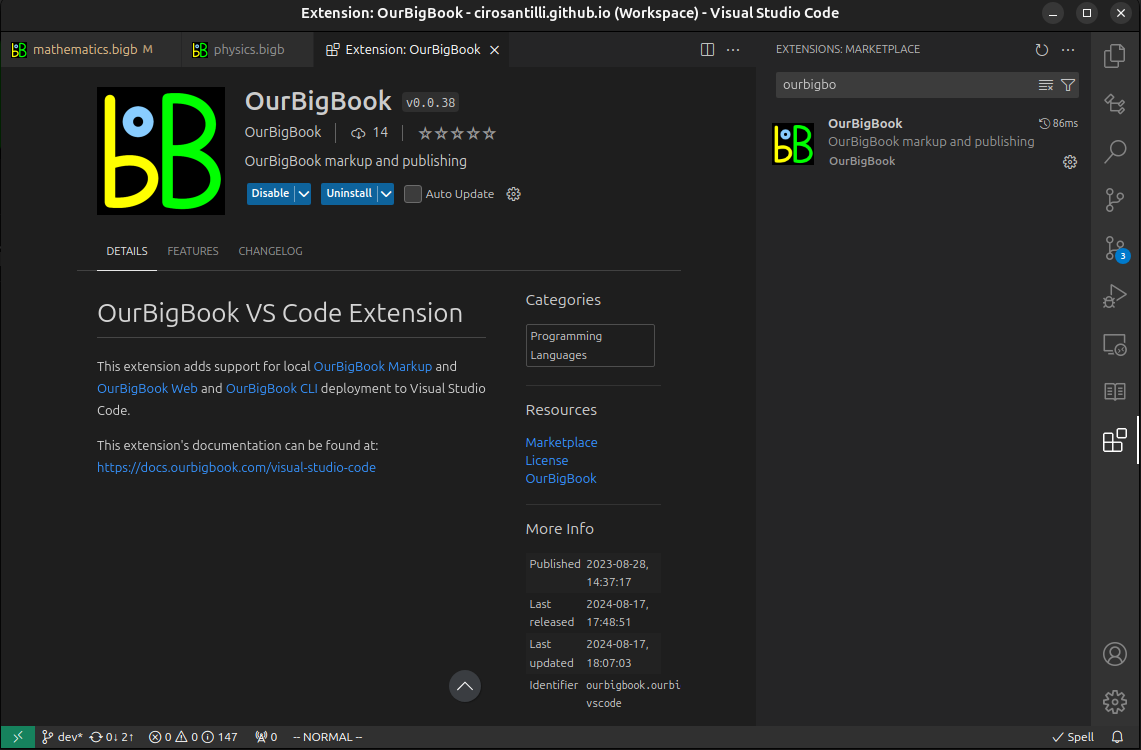
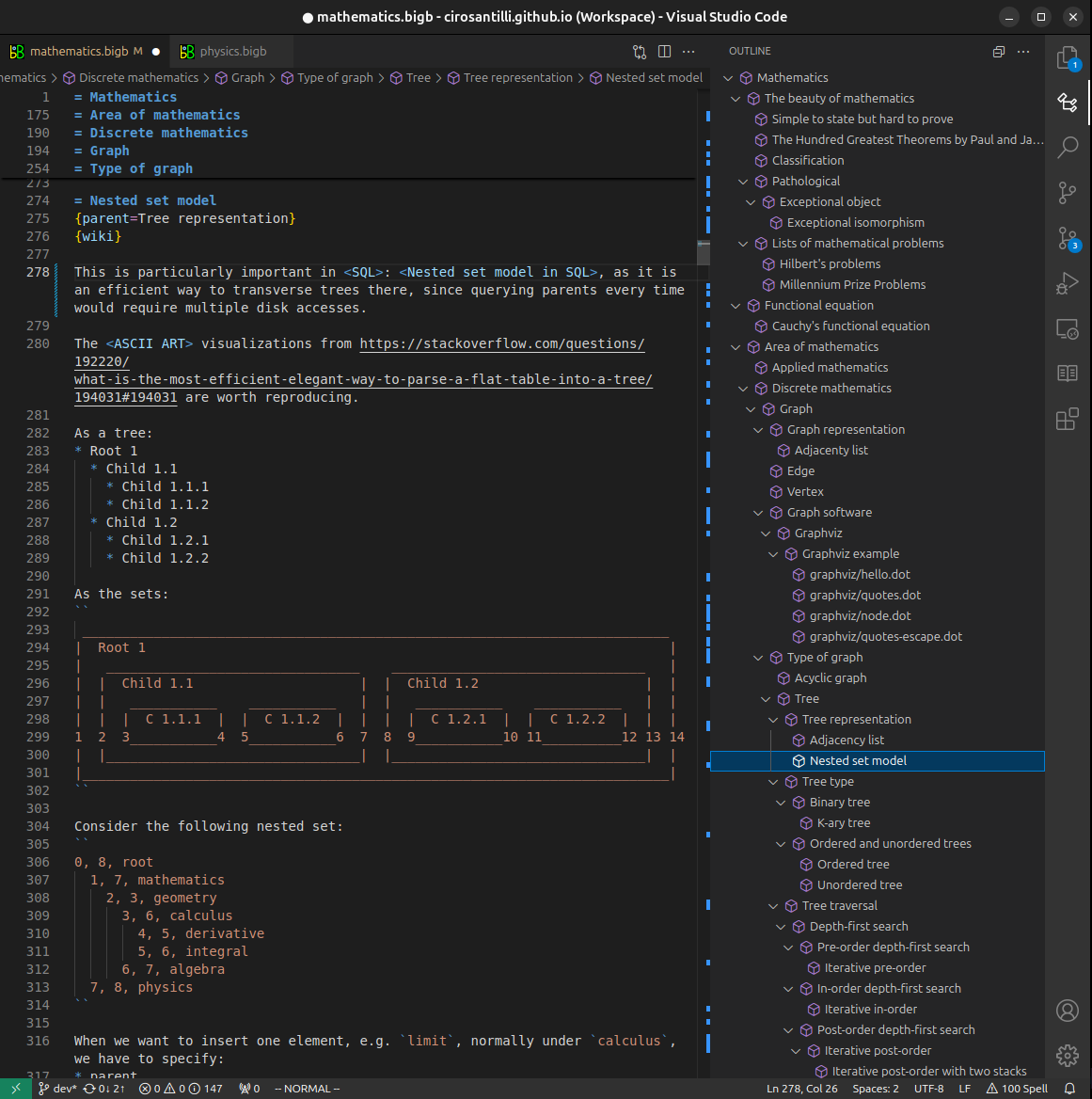
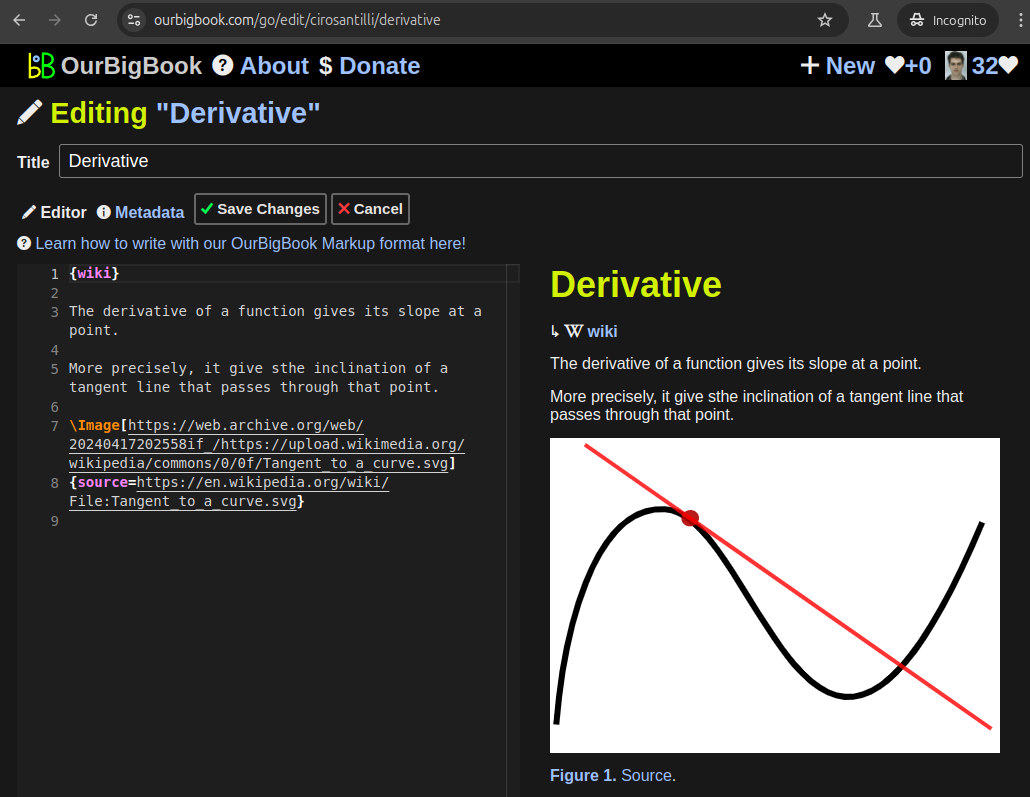
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact