An acoustic waveguide is a structure that confines and guides acoustic waves, primarily sound waves, in specific directions, much like an optical waveguide confines light. These waveguides can be made from various materials and can take various forms, including solid, liquid, or gaseous mediums. The primary purpose of an acoustic waveguide is to control the propagation of sound, allowing it to travel efficiently from one point to another while minimizing loss of energy due to scattering or absorption.
Colin Camerer is an influential American behavioral economist and a key figure in the field of experimental economics. He is known for his work on the intersection of game theory, psychology, and economics, particularly how human behavior deviates from traditional economic models that assume rational decision-making. Camerer has conducted extensive research on concepts such as bounded rationality, strategic interactions in games, and the implications of cognitive biases on economic decision-making.
Arthur J. Robson is a notable figure in the field of economics, particularly known for his contributions to evolutionary economics and resource economics. He has conducted research on topics such as the dynamics of economic systems, the role of innovation, and the interplay between ecological and economic systems. Robson's work often emphasizes the importance of understanding natural processes and human behavior in shaping economic outcomes.
David Schmeidler is an influential figure known for his work in economics, particularly in the fields of decision theory and game theory. He is recognized for contributions to the understanding of choice under uncertainty and the development of models that explain how individuals make decisions based on their preferences. His work often intersects with behavioral economics, and he is known for exploring foundational issues in economic theory.
Drew Fudenberg is an American economist known for his work in game theory, particularly in the areas of dynamic games, evolutionary game theory, and economic modeling. He has contributed significantly to the understanding of strategic interactions in economics and has published numerous influential papers on various topics, including repeated games and the evolution of social norms. Fudenberg has also co-authored textbooks that are widely used in the field of game theory and economics.
Hamidou Tembine is a researcher and professor known for his work in the fields of networking, systems, and AI. He has contributed significantly to areas such as wireless systems, network optimization, and machine learning applications within these domains. Tembine's research often involves mathematical modeling and analysis to address complex problems in communication networks, particularly in the context of improving the efficiency and performance of various network protocols and systems.
Eilon Solan is a prominent figure in the field of operations research and management science, particularly known for his work in game theory, optimization, and decision-making processes. He has made significant contributions to various areas, including bidding strategies, auction theory, and resource allocation.
Joel Sobel is a prominent American economist known for his contributions to game theory and related fields. He has worked extensively on topics such as strategic behavior, mechanism design, and the economics of information. Sobel's research often intersects with optimal decision-making and the functioning of markets. He is affiliated with the University of California, San Diego, where he has been involved in teaching and mentoring students in economics.
Marilda Sotomayor is a prominent figure in the field of mathematics, particularly known for her work in functional analysis, operator theory, and mathematical education. She has made significant contributions to the understanding of various mathematical concepts and is involved in research and teaching.
Georg Nöldeke (1832–1918) was a notable German scholar and philologist, particularly recognized for his contributions to the study of Islamic literature, including Arabic and Persian texts. He played a significant role in the analysis and interpretation of the Quran and Islamic historical literature. One of his well-known works is the "History of the Quran," in which he examined the evolution of the Quranic text and its historical context.
John Harsanyi (1920–2000) was a Hungarian-American economist and Nobel laureate known for his significant contributions to game theory, behavioral economics, and the theory of social choice. He was born in Hungary and faced persecution during World War II, which led him to emigrate to Australia, and later to the United States.
The term "Leighton relationship" generally refers to a concept in the field of mathematics, particularly in algebraic geometry and number theory, but it might also appear in other contexts. However, the most widely recognized usage pertains to the work of mathematician Leighton. In the context of algebraic geometry, it may describe relations or patterns between algebraic structures or could potentially relate to the behavior of certain mathematical properties under specific conditions.
Merrill M. Flood is best known for his contributions to the fields of mathematics, operations research, and game theory. He was an influential figure in the development of various mathematical concepts and methods, particularly those related to decision-making and optimization. He worked on problems such as the concept of bargaining and cooperative game theory, and he is recognized for his work on the Flood algorithm, which addresses network flows and related optimization problems.
Robert J. Elliott could refer to several individuals, as it is a relatively common name. However, one notable person is Robert J. Elliott, a prominent figure in the field of finance and risk management, particularly known for his contributions to the development of statistical methods and theoretical frameworks for financial applications. If you have a specific context in mind (such as finance, academia, literature, etc.
Werner Leinfellner is an Austrian philosopher known for his contributions to the fields of logic, language, and the philosophy of mathematics. He has been associated with various philosophical movements, including analytic philosophy and has worked on topics related to formal systems and the foundations of mathematics. Leinfellner's work often addresses the intersection of philosophy and mathematical logic, exploring the implications of logical systems for our understanding of language and reasoning.
A Coalition-proof Nash equilibrium (CPNE) is a solution concept in game theory that extends the traditional notion of Nash equilibrium to account for the possibility of coalition formation among players. In a standard Nash equilibrium, a strategy profile is stable if no single player can benefit by unilaterally changing their strategy, given the strategies of the other players. However, it does not consider the potential for groups of players to deviate together from the equilibrium, which can lead to different outcomes.
Suzanne Scotchmer (1948–2014) was an influential American economist recognized primarily for her contributions to the fields of public economics and intellectual property. She was a professor at the University of California, Berkeley, where she made significant contributions to the understanding of innovation, knowledge creation, and patent policy. Scotchmer's work often explored the intersection of economics and intellectual property, analyzing how different systems of protection for innovation affect economic growth and the dissemination of knowledge.
In topology, the **disjoint union** (also known as the coproduct in the category of topological spaces) is a way to construct a new topological space from a collection of topological spaces such that the new space captures the "disjointness" of the original spaces.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
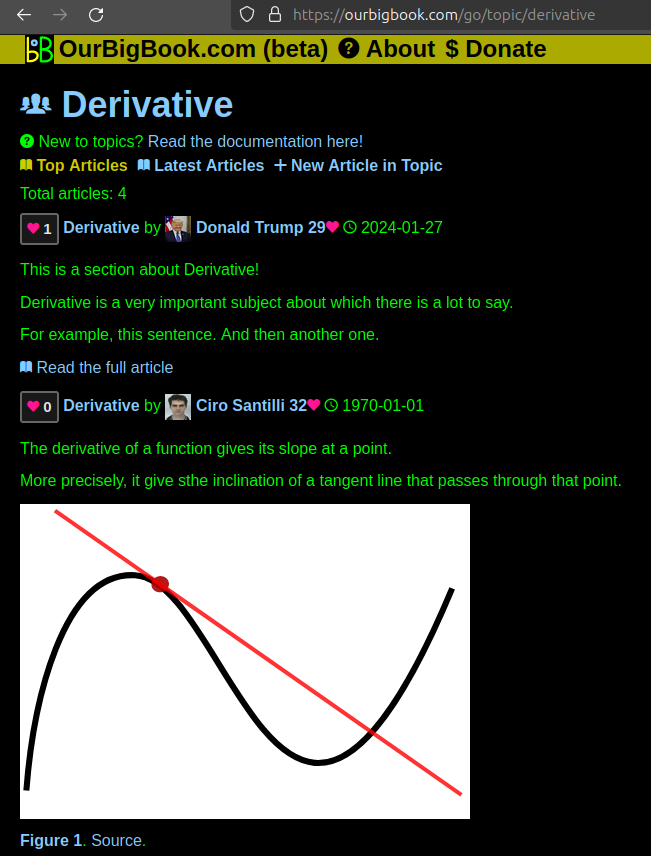
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
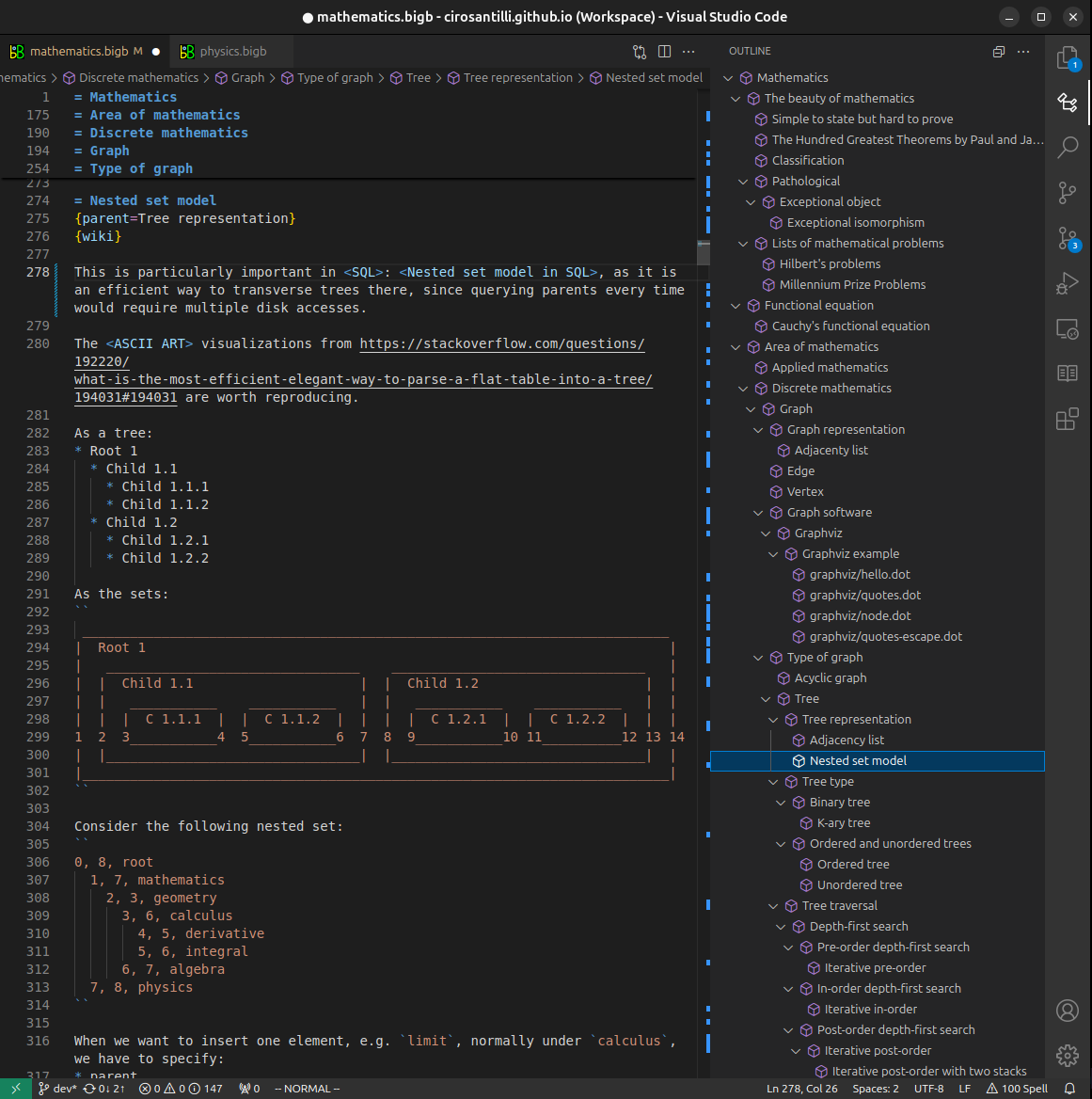
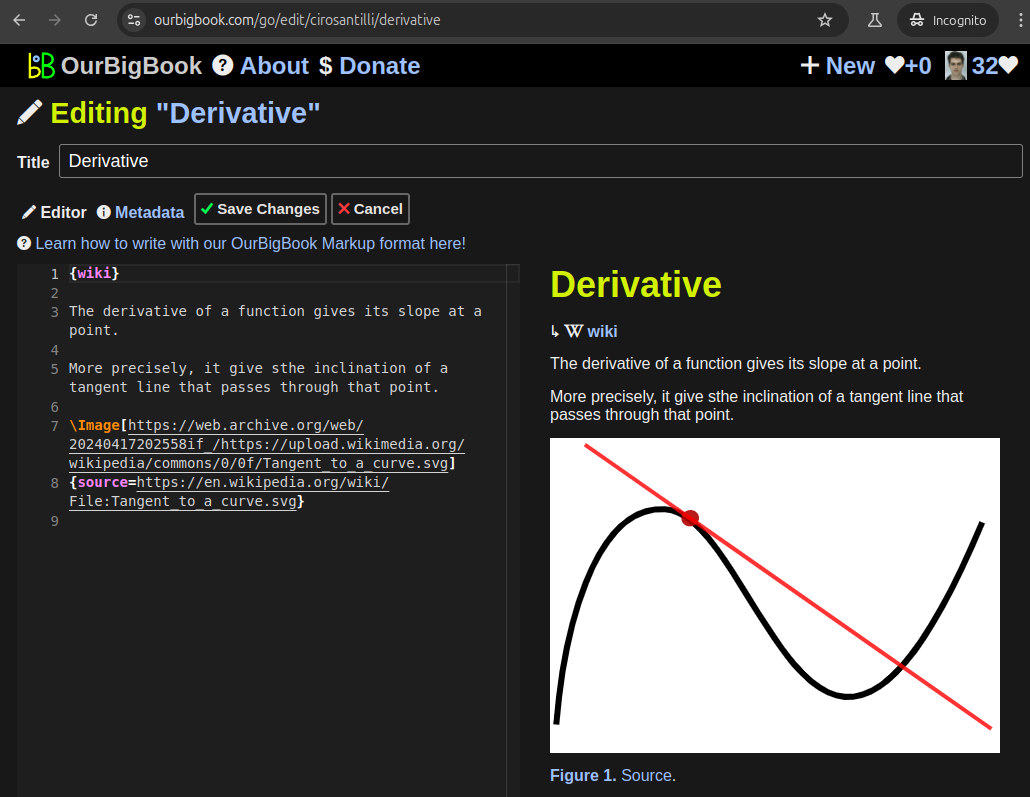
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact