Welcome to my home page!
RECOVER MONEY FROM ONLINE SCAM HIRE ADWARE RECOVERY SPECIALIST by  aliyaaitan1 0 Created 2025-02-24 Updated 2025-04-18
aliyaaitan1 0 Created 2025-02-24 Updated 2025-04-18
Email info: Adwarerecoveryspecialist@auctioneer.net
WhatsApp info:+1 (272) 332–8343
Website info: www.adwarerecoveryspecialist.com
Based in New York, I’ve always been familiar with the fast-paced and often volatile nature of financial markets. Navigating them has become second nature to me over the years, and I’ve had my share of both wins and losses. However, nothing prepared me for the emotional rollercoaster I experienced when I stumbled across a binary trading platform that promised incredibly high returns in a short period. The site looked legitimate, the testimonials seemed genuine, and the returns it promised were hard to ignore. Overcome by the potential for profit, I invested a substantial amount of Twenty Thousand Dollars. At first, everything seemed to be going as promised. I was getting regular updates on my supposed profits and felt confident that I had made the right move. However, as time went on, things began to feel off. The platform became increasingly difficult to access, and my attempts to withdraw funds were met with delays and excuses. It wasn’t long before I realized I had been scammed. The sinking feeling of losing everything was unbearable, and I found myself questioning how I could have been so careless. At my lowest point, when I was unsure if there was any way to recover my money, I came across a post on Instagram recommending a service called ADWARE RECOVERY SPECIALIST. At first, I was skeptical. After all, I had just been burned by a fraudulent platform, so the idea of trusting another service seemed risky. But the post highlighted their professionalism, quick action, and, most importantly, a track record of successfully helping people in similar situations. After some research and a few reassuring conversations, I decided to give them a shot. ADWARE RECOVERY SPECIALIST team was prompt, organized, and transparent in their approach. They understood the urgency of my situation and set to work immediately. Their professionalism reassured me that I was in good hands. Within just a few weeks, I received the news I had been hoping for my full investment of Twenty Thousand Dollars. was recovered. The relief I felt was indescribable. Not only had I regained my funds, but I also regained my faith in the process of recovery. I can’t thank ADWARE RECOVERY SPECIALIST enough for their dedication and expertise. Thanks to them, I’m now more cautious when navigating the online financial landscape, but I’m also grateful that there are services out there that genuinely care about helping people like me. If you ever find yourself in a similar situation, I highly recommend reaching out to them. They truly are a success story.
Welcome to my home page!
HIRE THE MOST EXPERIENCE CRYPTO SCAM RECOVERY EXPERT iFORCE HACKER RECOVERY by  Frank Patton 1 Created 2025-02-24 Updated 2025-04-18
Frank Patton 1 Created 2025-02-24 Updated 2025-04-18
I understand how frustrating it can be to invest online and end up losing money. I went through a similar experience when I invested in a binary option and was scammed out of 30,000 worth of BTC. It was a devastating loss, and I felt hopeless. Fortunately, I learned about iFORCE HACKER RECOVERY through a trusted friend who highly recommended them.At first, I was afraid, but I decided to give them a chance. To my amazement, they were able to recover my lost BTC within just 57 hours. I am incredibly grateful for their assistance and professionalism throughout the process. iFORCE HACKER RECOVERY truly saved me from a huge financial loss, and I can't thank them enough for their help.
Website; www. iforcehackersrecovery. com
Website; www. iforcehackersrecovery. com
Call/Text-whatsapp; +1 (240) 8033 (706)
Welcome to my home page!
Many East Asians, notably Chinese immigrants, choose to adopt a Western name pseudonym to make it easier for Western people who don't speak the language to call them and remember their name.
Cowards! Ciro Santilli would much rather just torture foreigners into learning his language. But fair play.
More interestingly however, some of the names chosen are not typical names, and some end up being very cute or mildly funny. Perhaps it is partly linked to given names are getting weirder.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
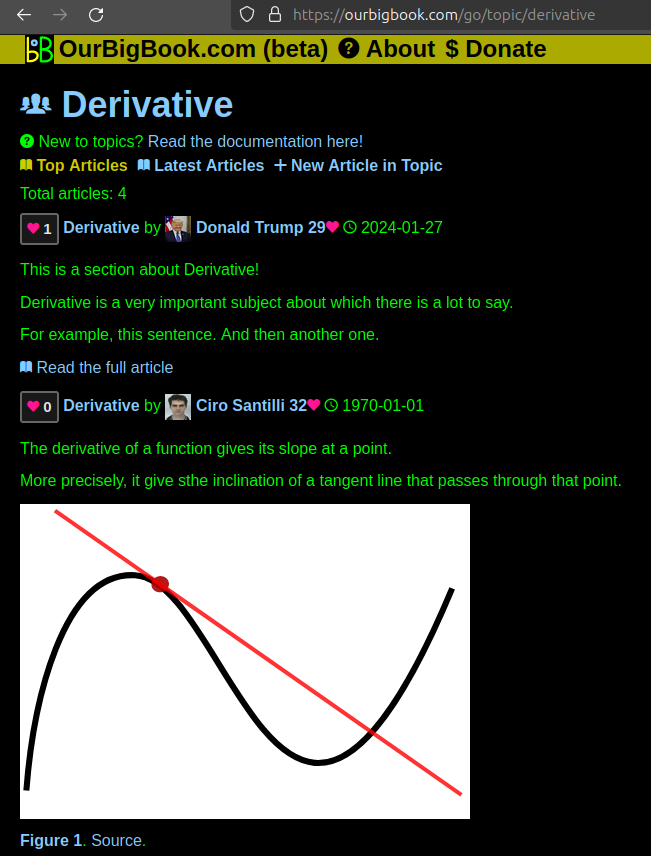
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
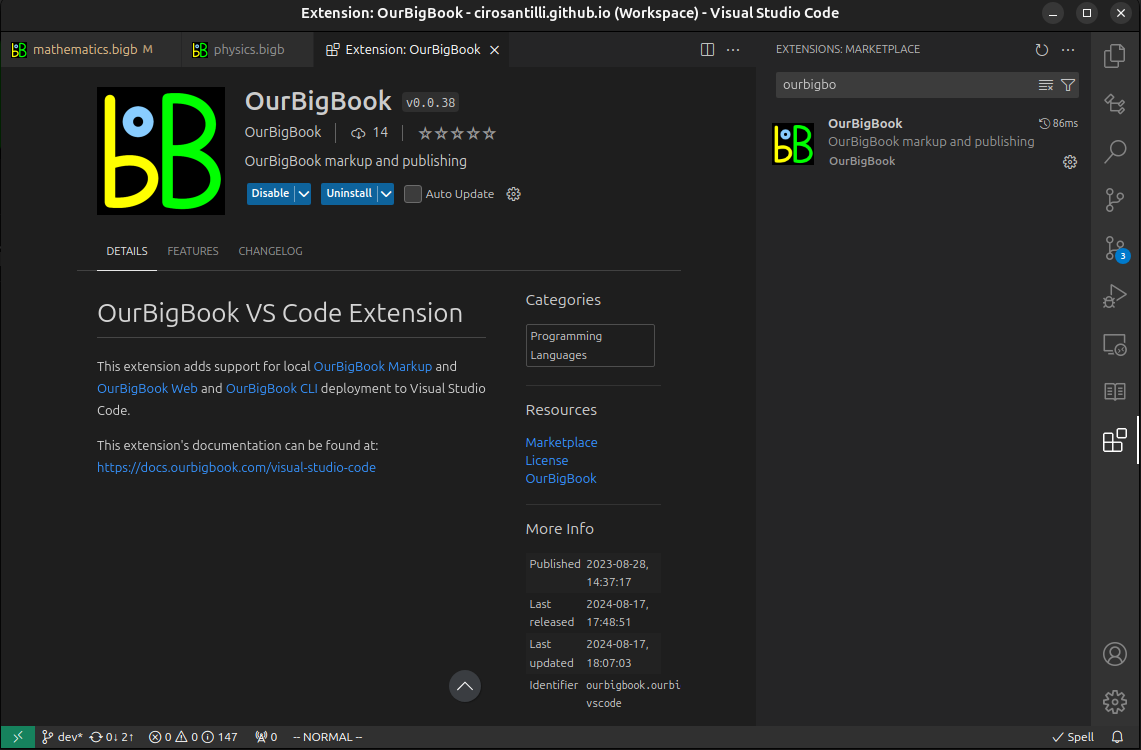
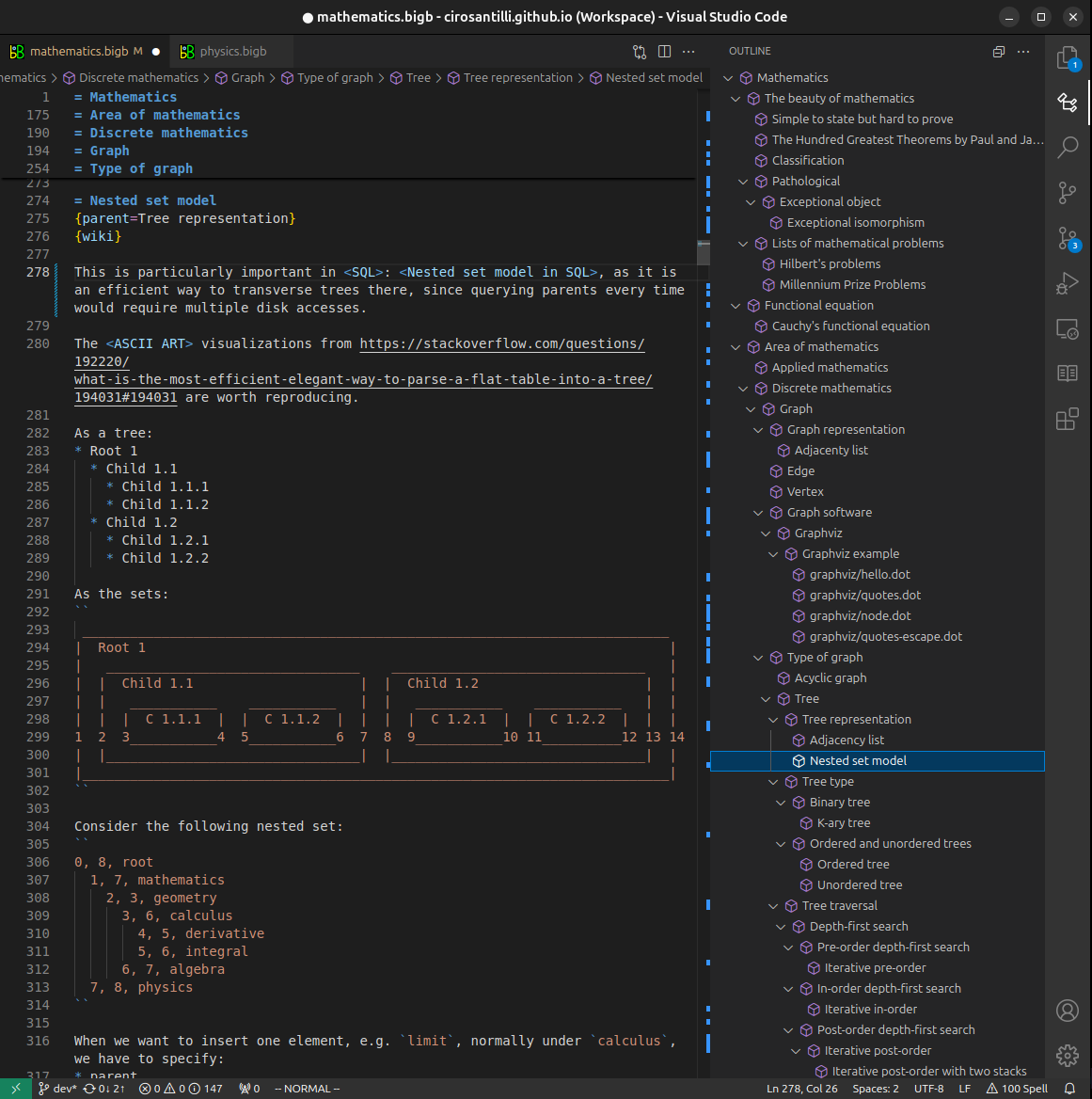
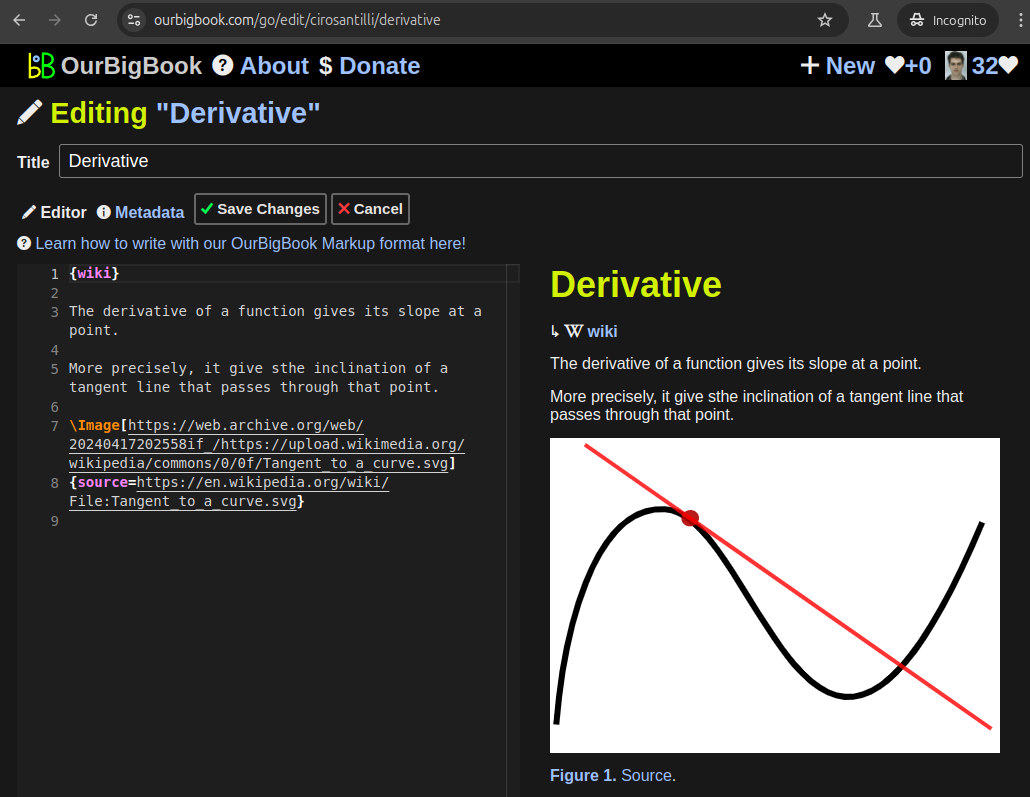
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact