Egyptian mathematicians refer to the mathematicians from ancient Egypt who made significant contributions to mathematics, particularly from around 3000 BCE to 300 CE. The ancient Egyptians developed a practical approach to mathematics that was closely tied to their needs in agriculture, trade, and engineering, especially in the construction of monumental architecture such as pyramids and temples.
As of my last knowledge update in October 2021, "Matheass" does not appear to be a widely recognized term, entity, or concept in mathematics, science, or popular culture. It's possible it could refer to a new concept, product, or specific niche community that has developed or gained popularity since then.
Mathspace is an educational platform designed to help students learn and practice mathematics through an interactive and adaptive online environment. The platform typically includes features such as step-by-step problem-solving, personalized learning paths, and real-time feedback, allowing students to work at their own pace and receive guidance as they tackle various math concepts. Mathspace aims to make math more engaging and accessible, often integrating multimedia resources, interactive exercises, and assessments.
NetTutor is an online tutoring service designed to provide students with access to academic support in a variety of subjects. It typically offers features such as live tutoring sessions with qualified tutors, on-demand homework help, and resources for specific subjects like math, science, writing, and more. NetTutor is often integrated into educational institutions' platforms, allowing students to easily access tutoring services as part of their course resources.
Egyptian nuclear physicists are scientists and researchers from Egypt who specialize in the field of nuclear physics. This branch of physics focuses on understanding atomic nuclei, their constituents, and their interactions. Nuclear physicists in Egypt may engage in various research areas such as nuclear energy, nuclear medicine, radiation physics, and particle physics.
Sameera Moussa was an Egyptian nuclear physicist, renowned for her pioneering work in the field of nuclear science and its applications in medicine, particularly in cancer treatment. Born in 1917, she made significant contributions to the development of peaceful uses of atomic energy. Moussa was notable for her efforts to promote the use of nuclear technology for medical purposes in Egypt and the broader Arab world.
Electrical Discharge Machining (EDM) is a manufacturing process used to remove material from a workpiece by using electrical discharges (sparks). It is particularly effective for machining hard materials and complex shapes that may be difficult to achieve using traditional machining methods. ### Key Features of EDM: 1. **Process Mechanism**: - EDM works on the principle of anodic dissolution.
A High-Enthalpy Arc Heated Facility (HEAHF) is a specialized research and testing facility designed to simulate the extreme thermal and aerodynamic conditions that aerospace vehicles experience during hypersonic flight or reentry into the Earth's atmosphere. The fundamental principle behind such facilities is the use of an electric arc to generate high temperatures and enthalpy levels, allowing researchers to study material responses, thermal protection systems, and aerodynamic characteristics in conditions that closely resemble those encountered in real flight scenarios.
Piezoelectric materials are substances that exhibit the piezoelectric effect, which is the ability to generate an electric charge in response to applied mechanical stress. This phenomenon occurs in certain materials when they undergo deformation, causing a separation of positive and negative charges within the material, thus creating an electric field. Key features of piezoelectric materials include: 1. **Types of Materials**: Common piezoelectric materials include certain crystals (e.g., quartz, topaz), ceramics (e.g.
Compact intracloud discharge (CID) refers to a type of electrical discharge that occurs within a thunderstorm cloud, specifically between different regions of the cloud itself, rather than between the cloud and the ground or between separate clouds. These discharges are often shorter and more compact than typical cloud-to-ground lightning strikes. CIDs are characterized by their localized nature and can occur within the complex structure of the cloud, which consists of various charged regions.
The Dember effect refers to a phenomenon observed in psychophysics, particularly in the study of perception and attention. Named after the psychologist William Dember, this effect describes how the presence of a secondary task or stimulus can influence the performance on a primary task, often enhancing or impairing it.
Lightning activity levels are classifications or scales used to describe the frequency and intensity of lightning strikes in a particular area over a specific period. These activity levels can help meteorologists and researchers understand and predict thunderstorms and severe weather. Lightning activity levels can vary based on different factors, including: 1. **Frequency**: The number of lightning strikes within a given area over a defined time frame. 2. **Intensity**: The strength or energy of the lightning, which can affect the damage it may cause.
Mains hum, often referred to as "60 Hz hum" in the United States or "50 Hz hum" in many other countries, is an audible electrical noise that is typically associated with alternating current (AC) power supply systems. **Causes of Mains Hum:** 1. **Power Frequency:** The hum arises because of the frequency of the AC power supply. In the U.S.
The piezoelectric coefficient is a measure of the efficiency with which a material converts mechanical energy into electrical energy (and vice versa) through the piezoelectric effect. It quantifies the relationship between the mechanical stress applied to a piezoelectric material and the resulting electric charge (or voltage) generated.
An active circulator is a type of radio frequency (RF) component used in microwave and antenna systems to direct the flow of signals. Unlike passive circulators, which rely solely on passive components and ferrite materials to route RF signals, active circulators incorporate active components, like amplifiers, to enhance signal transmission and improve overall performance.
A Ring Main Unit (RMU) is a type of electrical equipment used in medium-voltage distribution networks. It is designed to distribute electrical power while providing a degree of protection, control, and isolation. Typically, RMUs are used in urban areas and industrial applications where space is limited and reliability is critical.
The electrical system of the International Space Station (ISS) is a complex network designed to provide power to all on-board systems, equipment, and experiments. Here are its key components and features: ### Power Generation 1. **Solar Arrays**: The primary source of power for the ISS comes from large solar arrays. There are eight solar arrays that convert sunlight into electricity, providing approximately 84 kilowatts of power.
An air gap in a magnetic context refers to the physical space or gap that exists between magnetic materials or components within an electromagnetic system. This gap is typically filled with air (or another non-magnetic medium) and can significantly affect the magnetic flux and performance of devices such as transformers, relays, magnetic switches, and electric motors.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
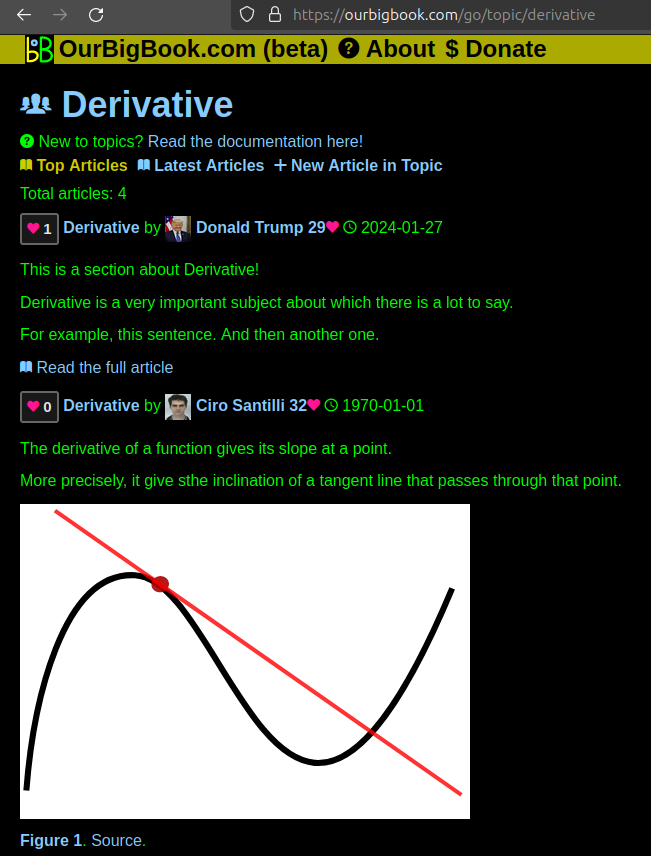
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
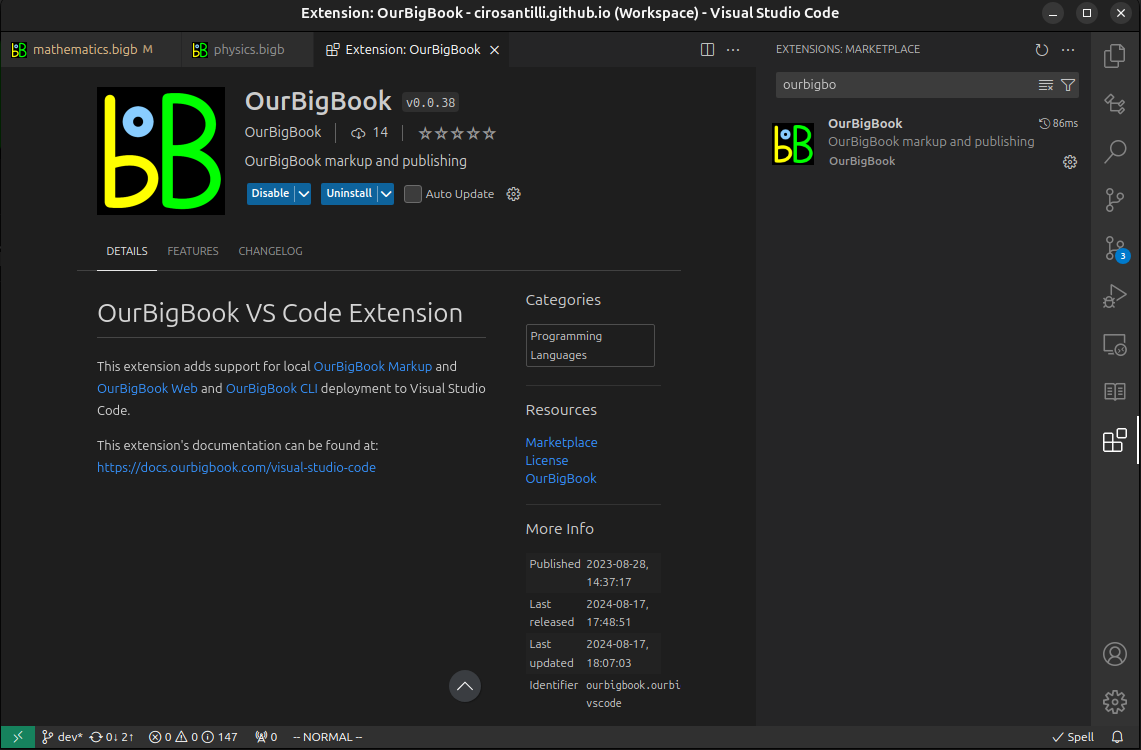
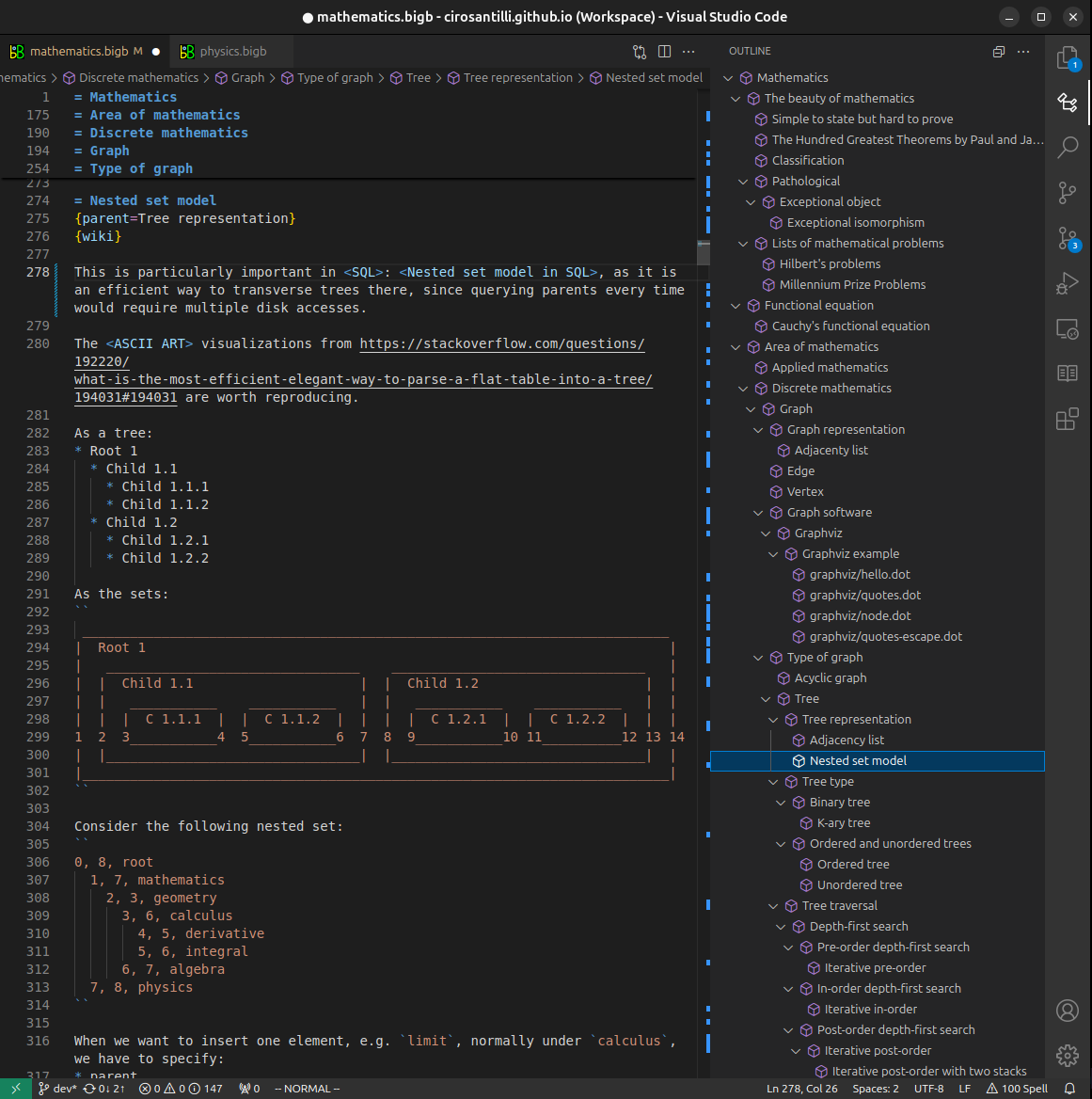
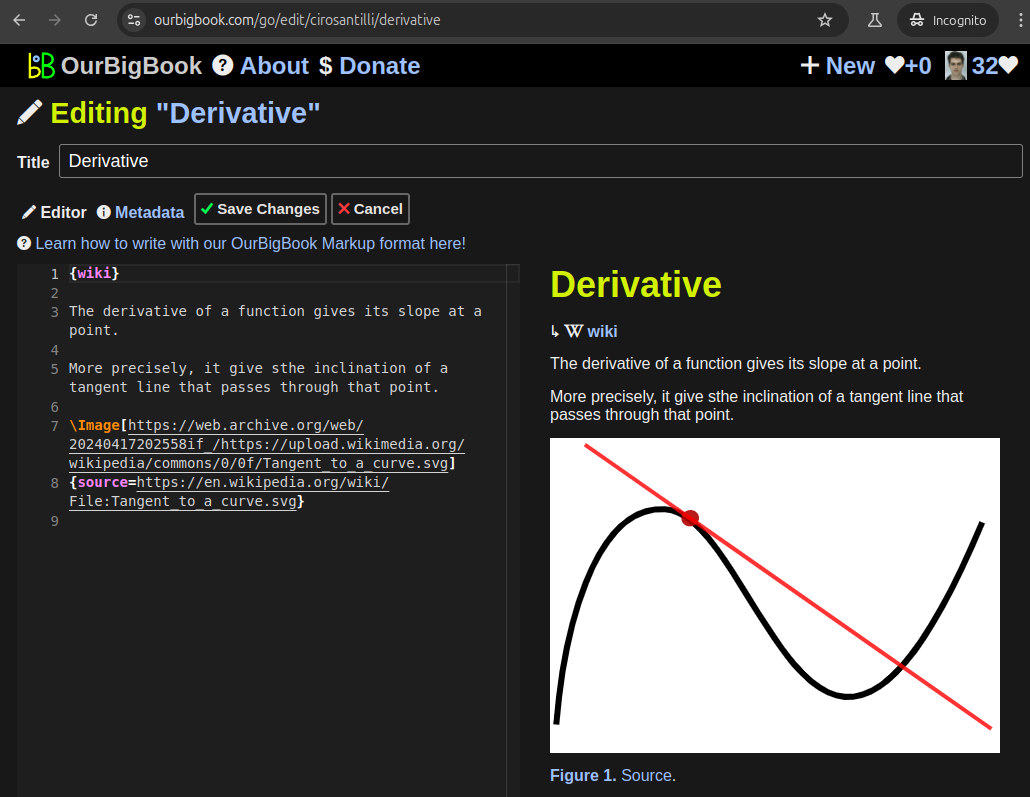
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact