Haakon Chevalier (1915–2009) was an American writer and academic known for his work as a novelist, playwright, and literary critic. He was also notable for his involvement in various literary and intellectual circles during the 20th century. Chevalier's writing often explored themes of identity, culture, and the human experience, reflecting his diverse background and the influences of his time.
Kenneth Nichols was an American engineer and military officer known for his significant contributions to the development of the atomic bomb during the Manhattan Project in World War II. He is particularly noted for overseeing the construction of the Los Alamos Laboratory in New Mexico, which was a key site for the research and development of nuclear weapons. After the war, Nichols held various leadership positions in the U.S. government and private sector, including working with the Atomic Energy Commission.
Eugen Merzbacher was a physicist known for his contributions to the field of quantum mechanics and for authoring educational materials. One of his notable works is the textbook "An Introduction to Quantum Mechanics," which is widely used in academic settings to teach the principles of quantum theory. His textbook is recognized for its clarity and depth, covering various topics such as wave-particle duality, quantum states, and the mathematical foundations of quantum mechanics.
Fay Ajzenberg-Selove (1930–2022) was a notable American physicist known for her contributions to the field of nuclear physics. She was particularly recognized for her research on the structure of atomic nuclei and the processes involved in nuclear reactions. Ajzenberg-Selove was also known for her work in promoting the role of women in science and for her efforts in advancing gender equity in the field of physics.
Oppenheimer is a lunar impact crater located on the Moon's surface. It is situated in the southern region of the Moon, near the boundary of the Mare Nectaris, a large basaltic plain formed by ancient volcanic activity. Oppenheimer is characterized by its roughly circular shape and has a rugged, eroded rim, which indicates that it has been subjected to impacts over a long period of time. The crater is named after J.
"Tryst" is an album by the American jazz artist and pianist Paul Bley, released in 1990. The album features a unique blend of jazz styles and showcases Bley's innovative approach to improvisation and composition. It typically includes a mix of original pieces and interpretations of jazz standards, highlighting Bley's interactive playing style and his ability to create atmospheric soundscapes. The album is well regarded among jazz enthusiasts for its artistic depth and interpretative versatility.
"Under the Influences" can refer to several different things depending on the context. Here are a few possibilities: 1. **Film or Television**: It might refer to a specific movie or episode of a TV show. There are several films and series that may have a similar name or theme related to influences, addiction, or substance use.
"Under the Streetlight" is a musical project and album by the American vocal group The 5th Dimension, released in 2017. The album is a collection of popular and traditional songs, blending elements of pop, R&B, and other genres. It features both original tracks and covers, showcasing the group's harmonies and musical versatility. The project may also refer to themes of nostalgia, community, and the stories that unfold under the metaphorical and literal lights of urban environments.
GABRG1 is a gene that encodes the gamma-aminobutyric acid (GABA) A receptor subunit gamma-1. The GABA A receptor is a major inhibitory neurotransmitter receptor in the central nervous system and plays a crucial role in mediating the effects of GABA, the primary inhibitory neurotransmitter in the brain. The GABRG1 subunit is one of several subunits that can assemble to form a functional GABA A receptor.
Canon Inc. is a Japanese multinational corporation that specializes in imaging and optical products, including cameras, camcorders, printers, and medical equipment. Founded in 1937 and headquartered in Tokyo, Canon is well-known for its innovation in photographic and imaging technology. The company initially gained recognition for its cameras, particularly film cameras, and later became a leader in the digital camera market, offering a wide range of products from compact point-and-shoot cameras to advanced DSLR and mirrorless systems.
"Werk 80" is the title of an album by the German band Atrocity, released in 1997. The album is notable for its unique approach, as it features covers of various songs from the 1980s, reinterpreted in a metal style. Atrocity is known for blending elements of death metal, industrial, and gothic music, and "Werk 80" showcases their ability to transform pop and rock hits from that era into heavier tracks.
"Ye Olde Space Bande" does not appear to correspond to any widely recognized concept, group, or piece of media up to my last training cut-off in October 2023. It may be a reference to a fictional or humorous concept blending elements of medieval or old-fashioned themes with a space setting, possibly created for entertainment purposes, such as in a video game, book, or online community.
Musical settings of poems by Arthur Rimbaud have been created by various composers across different musical genres. Rimbaud, a French poet known for his vivid imagery and innovative language, has inspired numerous musicians and composers to set his poems to music. Here are a few notable examples: 1. **"Il a neigé"** - This poem has been set to music by various composers, including the French songwriter and composer, Claude Debussy, who was known for his impressionistic style.
Musical settings of poems by Robert Louis Stevenson often highlight his lyrical and whimsical writing style. Stevenson, best known for his works such as "Treasure Island" and "Strange Case of Dr Jekyll and Mr Hyde," also wrote a number of poems that are beloved for their melodic qualities and imaginative themes, especially those found in his collection "A Child's Garden of Verses." Many composers have set Stevenson's poetry to music, tapping into the charm and innocence of his verses.
The Codex Sangallensis 381 is a notable manuscript of the Latin Bible, thought to have been produced in the early 9th century. It is housed in the Abbey Library of Saint Gall in Switzerland and is a significant source for the study of biblical texts and the history of the Bible in the medieval period. This codex is particularly interesting because it contains not just the biblical texts but also various glosses and commentaries that were added later.
The Leiden choirbooks, also known as the "Leiden Chorbücher," refer to a collection of choir books produced in the 16th century in the city of Leiden, Netherlands. These choirbooks are significant for their role in the history of music, particularly choral music, during the Renaissance period. The collection typically contains liturgical music, including masses, motets, and hymns, that were intended for use in church services.
Micrologus is a term that can refer to different subjects depending on the context. In the realm of music, Micrologus refers to an Italian early music ensemble known for performing Renaissance and medieval music. The group is recognized for its expertise in historical performance practices and its focus on authentic interpretations of ancient scores.
The Puy Manuscript, also known as the Puy Codex or the Codex of Puy, is a historical document that contains detailed records and information related to the customs, laws, and traditions of a specific community or region. Although there are several manuscripts referred to as "Puy Manuscript," one of the more notable references is to the legal document associated with the Puy de Fou, a historical theme park in France.
"Scolica enchiriadis" is a medieval treatise that is primarily known for its significance in the history of music theory. Written in the 9th century, it is attributed to Hucbald, a Benedictine monk and music theorist. The text serves as an introductory guide to music and discusses the concepts of pitch, melody, and harmony, as well as the notation and performance of music during that period.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
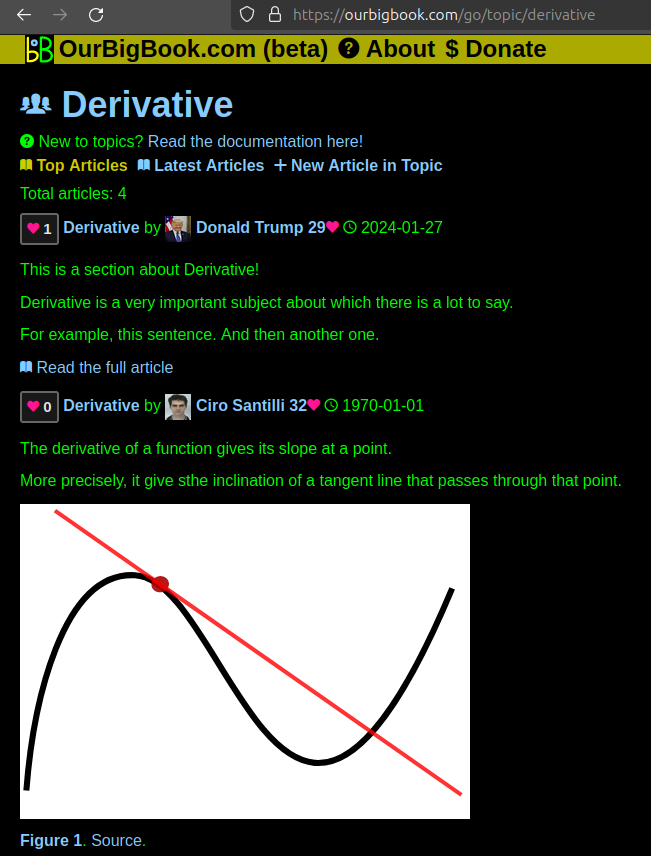
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
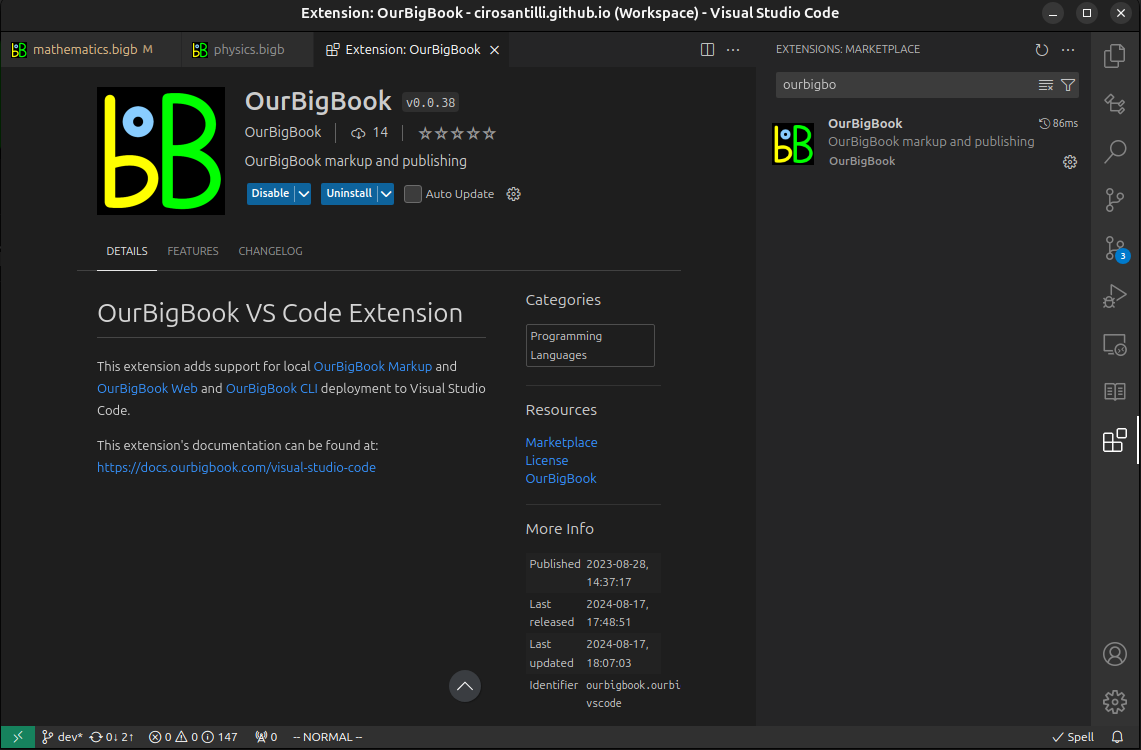
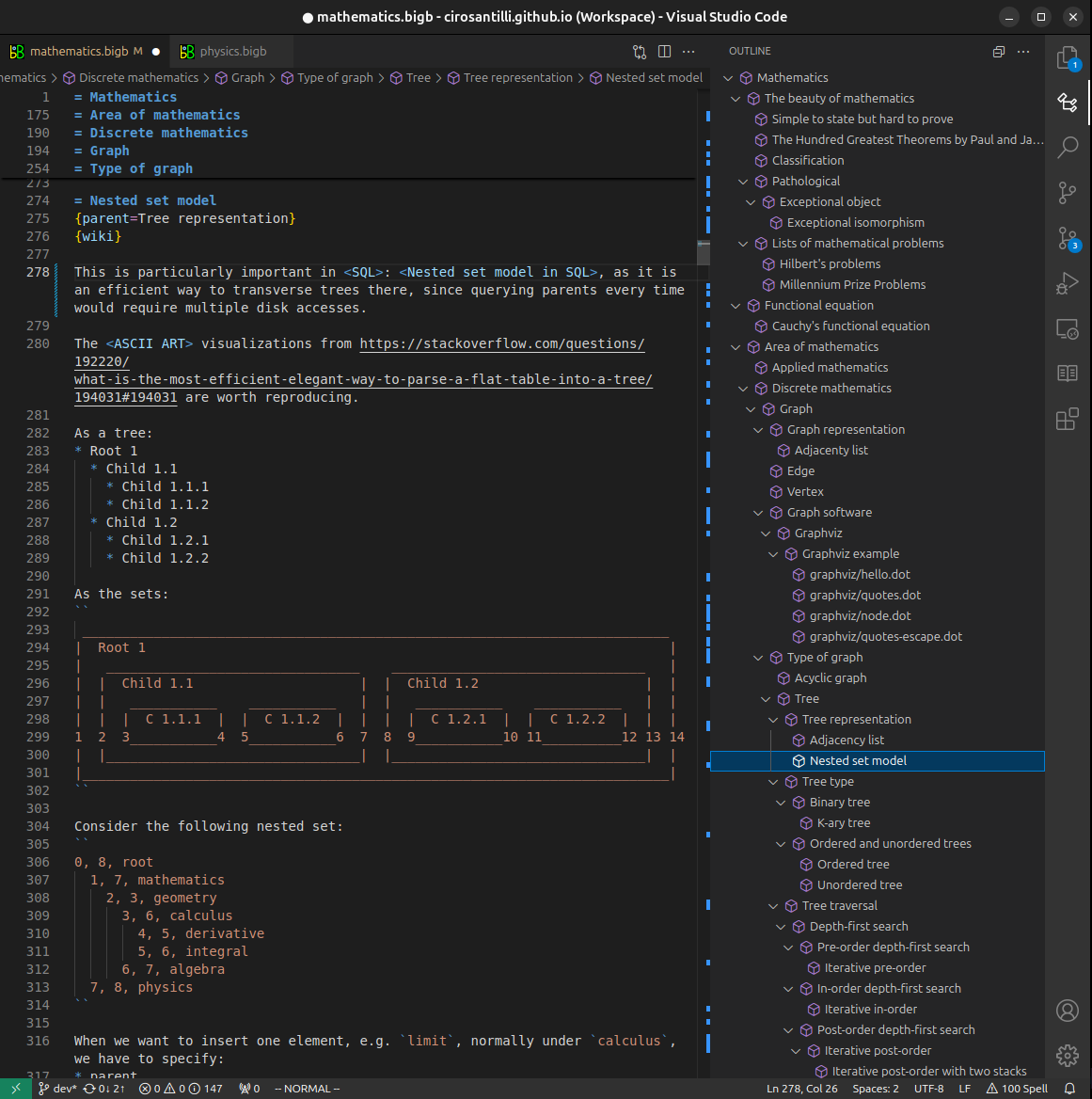
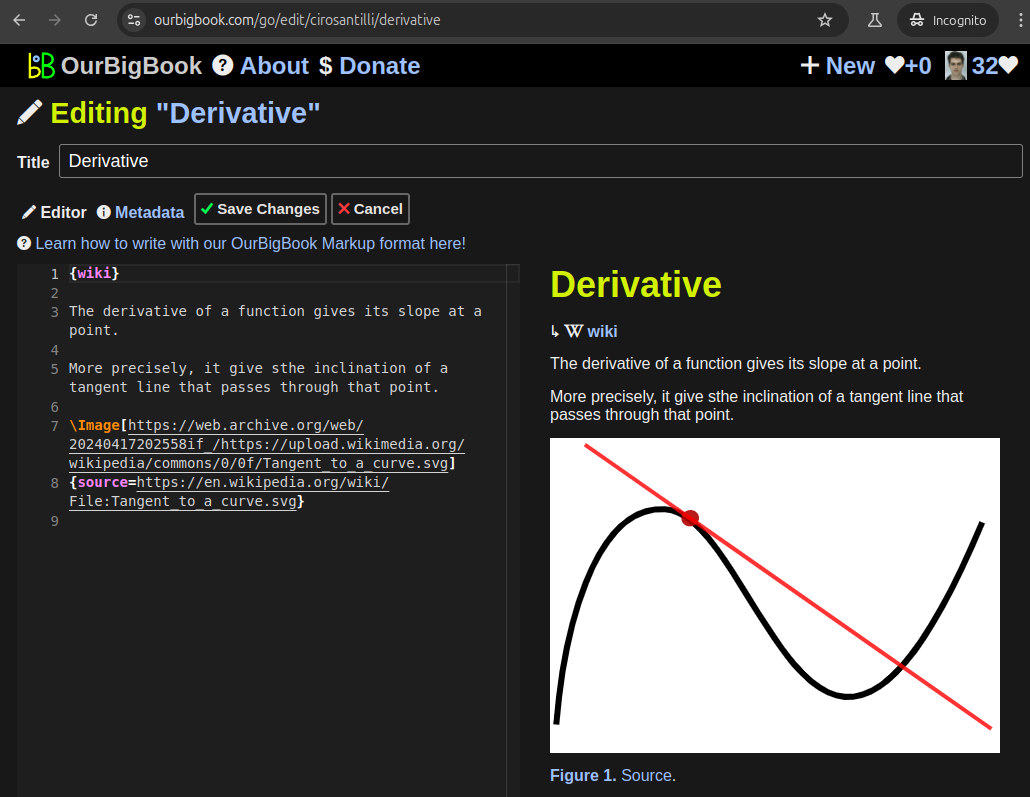
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact