Lichtner Seamount is an underwater volcanic feature located in the Pacific Ocean. Seamounts are essentially submarine mountains formed by volcanic activity. They can vary in height, shape, and size and often serve as important ecological habitats for various marine life. Lichtner Seamount, like other seamounts, may be of scientific interest for studies related to geology, marine biology, and oceanography.
The Makarov Basin is a region located in the Arctic Ocean, specifically within the East Siberian Sea. It is situated north of the Siberian coast and is bordered by the Severnaya Zemlya islands to the west and the New Siberian Islands to the east. The basin is characterized by its relatively deep waters, which are part of the complex underwater topography of the Arctic. The Makarov Basin is named after the Russian explorer and naval officer, Mikhail Makarov.
A Varifold is a mathematical concept used in differential geometry and geometric measure theory. It generalizes the notion of a manifold by allowing for more flexibility in the way that "sheets" of the object can intersect and overlap. Varifolds are typically used to study objects that may not have a well-defined smooth structure everywhere, such as irregular shapes, and they are particularly useful for analyzing geometric issues in a more robust way than traditional manifolds.
The Luzin \( N \) property is a concept from real analysis and functional analysis, particularly in the context of measurable functions. A function \( f: \mathbb{R} \to \mathbb{R} \) is said to have the Luzin \( N \) property if for every measurable set \( E \) of finite measure, the image \( f(E) \) is also a measurable set of finite measure.
Norman E. Gibbs is a mathematician known for his contributions to various fields, including statistics and mathematical modeling. However, without more context, it's difficult to provide a specific answer, as there may be multiple individuals with that name or various contexts in which he might be known.
Korean creation narratives are traditional stories or myths that explain the origins of Korea, its people, and the world. These narratives often incorporate elements of mythology, folklore, and history, reflecting the cultural and spiritual beliefs of the Korean people. One of the most well-known creation myths in Korea is the story of **Dangun** (or Dan-gun), who is considered the legendary founder of the Gojoseon kingdom, often cited as the first Korean state.
Dan Farmer is a well-known figure in the field of computer security, particularly recognized for his work in network security and vulnerability assessment. He is notable for co-developing the popular security tool "SATAN" (Security Administrator Tool for Analyzing Networks) in the early 1990s, which was one of the first tools designed to automate the process of network vulnerability scanning. Farmer has also contributed to various discussions and projects related to computer security, risk management, and cybersecurity best practices.
An "interfering thread nut" generally refers to a specific type of fastener that is designed to create a secure fit with a threaded fastener, such as a bolt or screw, by having threads that are slightly larger in diameter than the corresponding external threads. The term "interfering" indicates that there is an intentional design feature where the internal threads of the nut are interfered with the outer threads of the bolt.
Cyclone Martin was a significant tropical cyclone that occurred in the southwest Indian Ocean during the 1999 cyclone season. It formed in early March 1999 and was notable for impacting several islands in the region, including Madagascar and Mauritius. It reached a peak intensity of around 120 km/h (75 mph), classified as a moderate tropical storm on the Saffir-Simpson scale.
MultiDark is a large cosmological simulation project designed to study the formation and evolution of large-scale structures in the universe, particularly with respect to dark matter and its influence on galaxy formation. The simulation includes a variety of cosmological parameters and uses advanced computational techniques to model and analyze the behavior of dark matter, baryonic matter, and the effects of gravity over vast cosmic scales.
Indigenous Aryanism is a concept that primarily refers to a nationalist ideology which posits that the Indigenous peoples of India, particularly those who are part of the Hindu tradition, are descendants of the ancient Aryans. Proponents of this ideology often claim that these Aryans were not foreign invaders but rather indigenous to the Indian subcontinent.
George Varghese can refer to a few notable individuals, depending on the context. One prominent figure is a computer scientist and professor known for his work in networking and systems. He has contributed significantly to the fields of computer networking and algorithms, particularly in areas like network management and routing. Another possibility is that George Varghese is a common name in various cultures, particularly in India and among the Indian diaspora.
DEXRON is a trademarked name for a series of automotive transmission fluids developed by General Motors (GM). The DEXRON specification was introduced in the 1960s for automatic transmissions and has undergone several iterations to adapt to evolving technology and performance requirements in automotive applications.
Paleohydrology is the study of the historical patterns and dynamics of water systems through geological time. It focuses on understanding the behavior of rivers, lakes, glaciers, and groundwater in the past by examining sedimentary records, geological formations, and other indirect indicators of hydrological conditions. Researchers in paleohydrology use various techniques, including: 1. **Sediment Analysis**: Examining sediment layers to infer past water levels, flow rates, and environments.
Business Process Validation (BPV) is a systematic approach used to ensure that business processes are functioning as intended, meeting specified requirements, and achieving desired outcomes. This validation is particularly important in regulated industries, such as pharmaceuticals, biotechnology, and healthcare, but it is also relevant in various sectors seeking to optimize their operations. Key components of Business Process Validation include: 1. **Definition of Processes**: Clearly defining and documenting the business processes that need validation, including inputs, outputs, roles, and responsibilities.
"Discoveries" by Emil Ernst is a work that explores themes related to science, philosophy, and the quest for knowledge. The book presents insights into various scientific discoveries and their implications for understanding the world and humanity's place within it. Emil Ernst, as an author, tends to blend concepts from different disciplines, emphasizing the interconnectedness of knowledge and the impact of discoveries on society.
Garth Gibson is a notable figure in the field of computer science, particularly recognized for his contributions to computer systems and networking. He is known for his work on various topics, including storage systems and file systems. One of his most significant contributions is in the development of the Andrew File System (AFS), which has had a substantial impact on distributed file systems.
Gary T. Leavens is a prominent computer scientist known for his work in the fields of software engineering, programming languages, and formal methods. He has made significant contributions, particularly in the areas of specification and verification of software systems, as well as research on programming language design and semantics. Leavens is also known for his work on the Java Modeling Language (JML), which is used for specifying the behavior of Java classes and methods.
Gerth Stølting Brodal is a Norwegian biologist known for his work in the field of marine biology, particularly regarding oceanography and fish research. He has contributed to various studies related to the ecology and behavior of marine organisms in their natural environments. His research often emphasizes the importance of understanding marine ecosystems in the face of environmental changes and human impacts.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
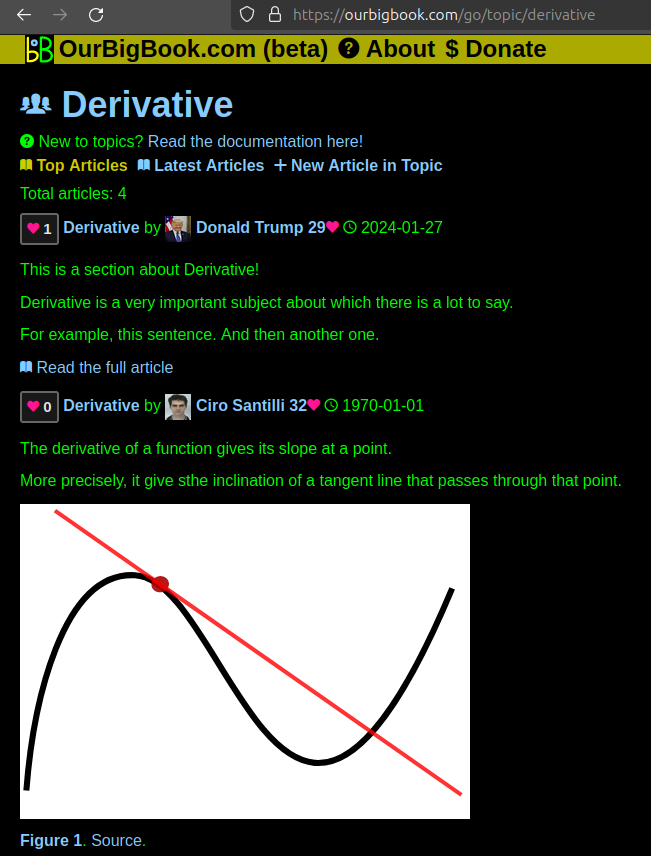
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
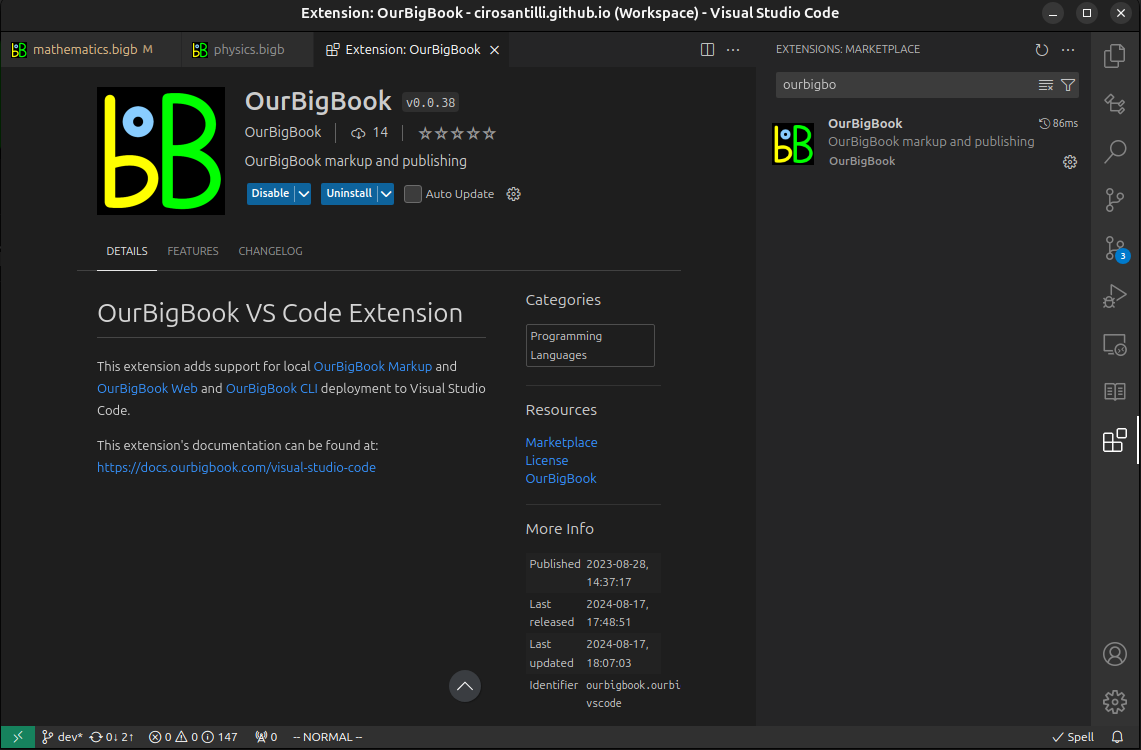
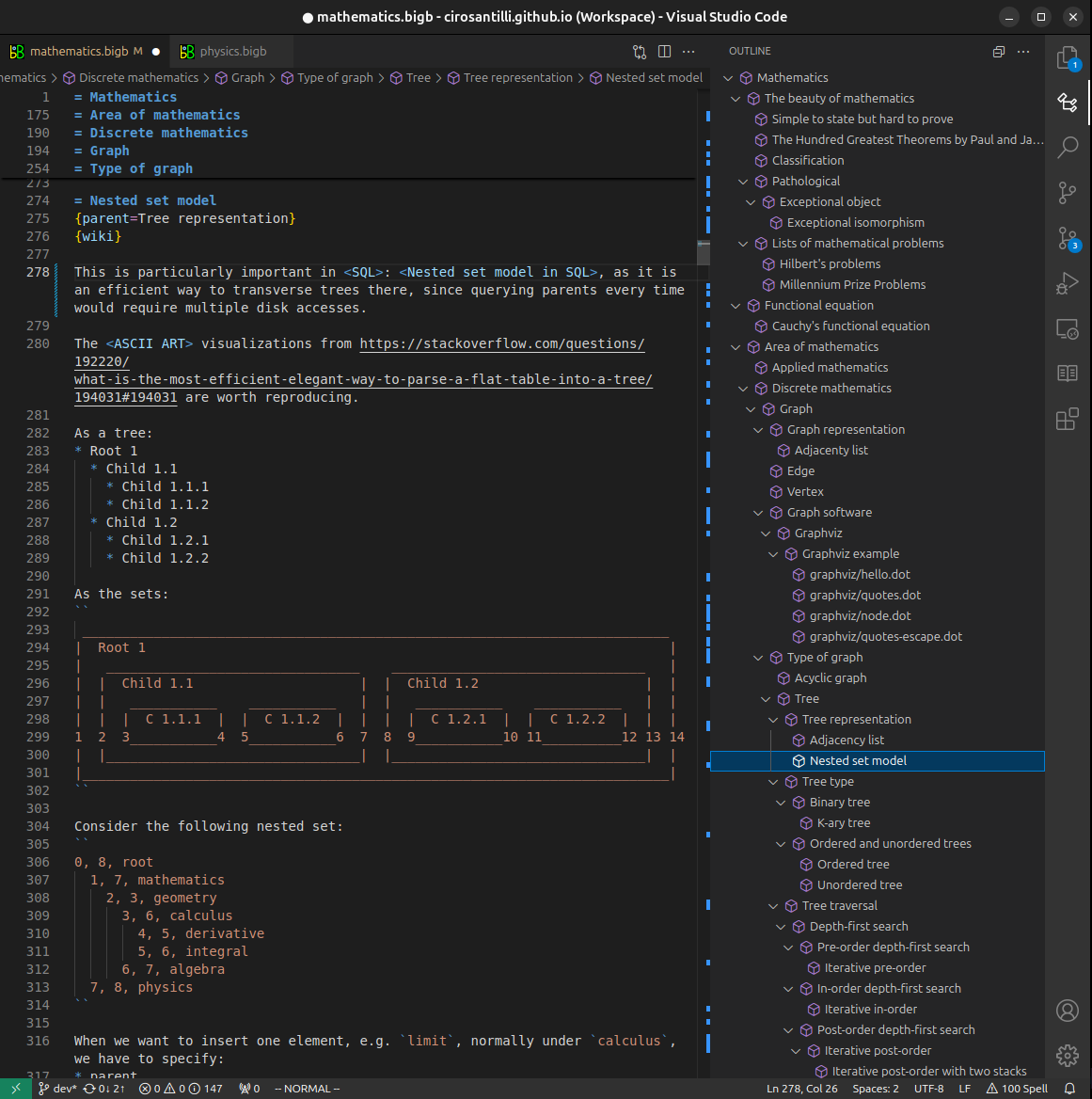
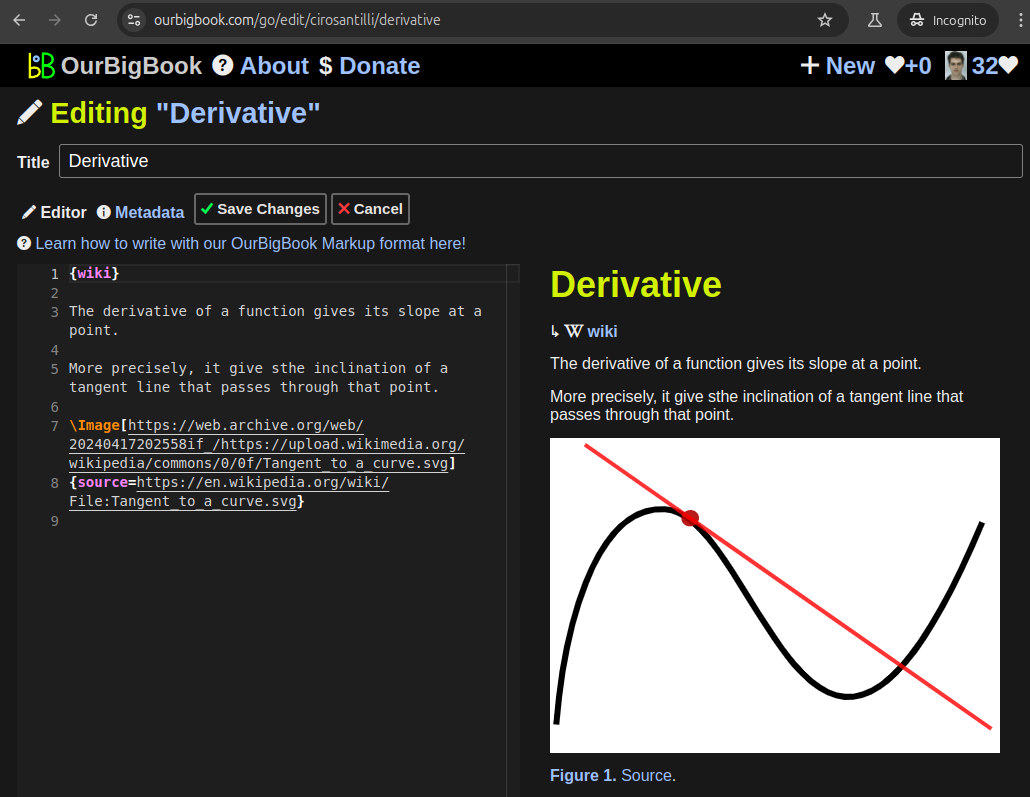
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact