Dmitri Mendeleev was a Russian chemist best known for developing the Periodic Table of Elements. Born on February 8, 1834, in Siberia, he made significant contributions to the field of chemistry, most notably in 1869 when he published his version of the periodic table. Mendeleev's table organized the known elements by their atomic mass and demonstrated that elements with similar properties appeared at regular intervals, a concept known as periodicity.
Dmitry Tolstoy can refer to a couple of notable figures, but it is most commonly associated with Dmitry Tolstoy (1818–1885), a prominent Russian statesman, and educator. He served in various governmental roles during the reign of Tsar Alexander II and was an important figure in Russian educational reform, advocating for more progressive educational policies.
Macarius Bulgakov (also known as Mikhail Bulgakov) is a notable figure from the early 20th century, particularly in the context of Russian literature and religious thought. However, it seems you might be referring to Macarius of Moscow, a significant saint in the Russian Orthodox Church, or Macarius Bulgakov, a prominent Orthodox theologian and bishop in the 20th century.
"Bending" can refer to different concepts depending on the context. Here are a few interpretations: 1. **Physical Bending**: In a mechanical context, bending refers to the deformation of a material when a force is applied. This can occur in various materials such as metals, plastics, and wood, and is often discussed in engineering and physics with respect to stress and strain.
"Freguesia" is a term used primarily in Portugal and some of its former colonies to refer to a civil parish or administrative district. Historically, it originated as a religious designation associated with a church or religious community, but over time it has evolved into a secular administrative unit. In Portugal, a freguesia is typically the smallest administrative division, serving various local governance functions, such as managing local services and community needs.
The Center for Advanced Study in the Behavioral Sciences (CASBS) is a research institute located at Stanford University, California. Established in 1954, CASBS is dedicated to advancing scholarly research in the behavioral and social sciences. It offers a collaborative environment for researchers, scholars, and practitioners from various disciplines to work together on complex social issues, engaging in intensive research, discussion, and reflection.
"Some Institutes for Advanced Study" typically refers to a number of prestigious institutions dedicated to research and scholarship in various disciplines, including the sciences, humanities, and social sciences. These institutes often aim to foster interdisciplinary collaboration and provide a conducive environment for scholars to engage in deep inquiry and innovative research.
The Oberwolfach Research Institute for Mathematics (Oberwolfacher Institute für Mathematische Forschung) is a renowned research institution located in Oberwolfach, Germany. Established in 1954, the institute is known for its focus on advanced research in mathematics and serves as a meeting place for mathematicians from around the world. The institute organizes workshops, which are typically small and focused, bringing together researchers working on specific mathematical topics.
The Statistical and Applied Mathematical Sciences Institute (SAMSI) is a research institute located in the United States that focuses on advancing the fields of statistical and applied mathematical sciences. Established to promote collaboration between statisticians, applied mathematicians, and researchers from various fields, SAMSI fosters interdisciplinary research in areas where statistics and applied mathematics intersect with other disciplines, such as biology, finance, social sciences, and engineering.
The JOLTS report, which stands for the Job Openings and Labor Turnover Survey, is a monthly report published by the U.S. Bureau of Labor Statistics (BLS). The primary purpose of the JOLTS report is to provide insights into labor market dynamics by tracking job openings, hires, separations, and turnover rates in the U.S. economy.
The NTNU University Museum, also known as the Norwegian University of Science and Technology (NTNU) University Museum, is a museum located in Trondheim, Norway. It is part of NTNU and serves as a center for research, education, and public engagement in various fields, including natural history, archaeology, and cultural heritage. The museum houses a diverse collection of artifacts, specimens, and exhibitions that reflect the cultural and natural history of Norway and the broader world.
The Lao Statistics Bureau (LSB) is a government agency in Laos responsible for collecting, analyzing, and disseminating statistical data to support national development and policy-making. It plays a crucial role in ensuring that accurate and timely statistics are available to inform government decisions, facilitate economic planning, and promote research and development. The LSB conducts various surveys, censuses, and data collection initiatives, including the national population and housing census, agricultural surveys, and economic indicators.
The Basque Statistics Office, known as "Eustat" (in Basque, "Euskal Estatistika Erakundea"), is the official statistical agency of the Basque Autonomous Community in Spain. It is responsible for collecting, analyzing, and disseminating statistical information about various aspects of the Basque Country, including demographics, economy, society, and the environment.
The Bureau of Transportation Statistics (BTS) is an agency within the United States Department of Transportation (USDOT). Its primary role is to collect, analyze, and disseminate data related to transportation in the U.S. The mission of BTS includes providing statistical information that can be used to inform decision-making by policymakers, researchers, and the public regarding transportation policy and infrastructure.
The Ghana Statistical Service (GSS) is a governmental agency responsible for collecting, analyzing, and disseminating statistical data in Ghana. Its primary role is to provide reliable statistical information that can inform policy-making, planning, and decision-making processes across various sectors of the economy and society. Key functions of the GSS include: 1. **Census and Surveys**: Conducting national and local censuses, including the Population and Housing Census, as well as various thematic surveys (e.g.
The Dominion Bureau of Statistics (DBS) was the official statistical agency of Canada, established in 1918 to collect, compile, and publish statistical information relating to the country's economy, society, and demographics. Its main purpose was to provide reliable and comprehensive statistical data to support government decisions, policy-making, and economic planning.
The National Statistical Institute (NSI) of Bulgaria is the official governmental body responsible for collecting, processing, and disseminating statistical information about various aspects of the country's economy, society, and environment. Established in 1880, it plays a crucial role in providing reliable and relevant statistical data that supports decision-making processes for government, businesses, and researchers.
The National Statistics Institute (Instituto Nacional de Estadística, INE) is the primary institution responsible for collecting, analyzing, and disseminating statistical information in Spain. Established in 1945, the INE operates under the Ministry of Economic Affairs and Digital Transformation and is tasked with producing official statistics that inform policy-making and provide essential data about the country's economy, demographics, and society.
Statistics Austria (Statistik Austria) is the national statistical office of Austria, responsible for collecting, processing, and disseminating statistical information about the country. Established to support the development of evidence-based policies and decision-making, it provides a wide range of data on demographics, economics, social conditions, environment, and public health, among other areas.
Statistics New Zealand, also known as Tatauranga Aotearoa, is the government agency responsible for collecting, analyzing, and disseminating official statistical information in New Zealand. Its primary role is to provide reliable and relevant statistical information to support decision-making, policy formulation, and research in the country. This includes conducting national censuses, surveys, and other data collection initiatives covering various aspects such as population, economy, health, and social factors.
Pinned article: Introduction to the OurBigBook Project
Welcome to the OurBigBook Project! Our goal is to create the perfect publishing platform for STEM subjects, and get university-level students to write the best free STEM tutorials ever.
Everyone is welcome to create an account and play with the site: ourbigbook.com/go/register. We belive that students themselves can write amazing tutorials, but teachers are welcome too. You can write about anything you want, it doesn't have to be STEM or even educational. Silly test content is very welcome and you won't be penalized in any way. Just keep it legal!
Intro to OurBigBook
. Source. We have two killer features:
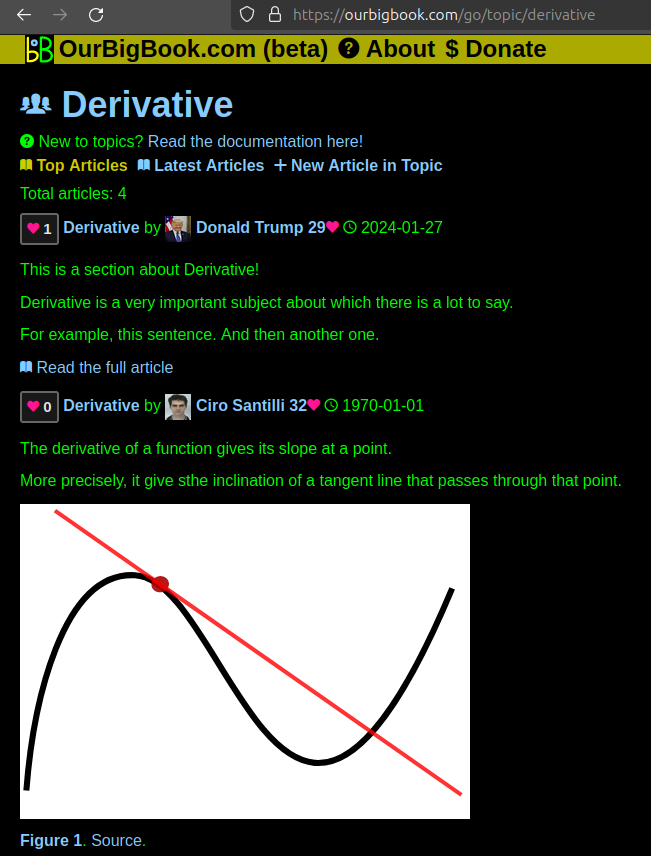
- topics: topics group articles by different users with the same title, e.g. here is the topic for the "Fundamental Theorem of Calculus" ourbigbook.com/go/topic/fundamental-theorem-of-calculusArticles of different users are sorted by upvote within each article page. This feature is a bit like:
- a Wikipedia where each user can have their own version of each article
- a Q&A website like Stack Overflow, where multiple people can give their views on a given topic, and the best ones are sorted by upvote. Except you don't need to wait for someone to ask first, and any topic goes, no matter how narrow or broad
This feature makes it possible for readers to find better explanations of any topic created by other writers. And it allows writers to create an explanation in a place that readers might actually find it.Figure 1. Screenshot of the "Derivative" topic page. View it live at: ourbigbook.com/go/topic/derivativeVideo 2. OurBigBook Web topics demo. Source. - local editing: you can store all your personal knowledge base content locally in a plaintext markup format that can be edited locally and published either:This way you can be sure that even if OurBigBook.com were to go down one day (which we have no plans to do as it is quite cheap to host!), your content will still be perfectly readable as a static site.
- to OurBigBook.com to get awesome multi-user features like topics and likes
- as HTML files to a static website, which you can host yourself for free on many external providers like GitHub Pages, and remain in full control
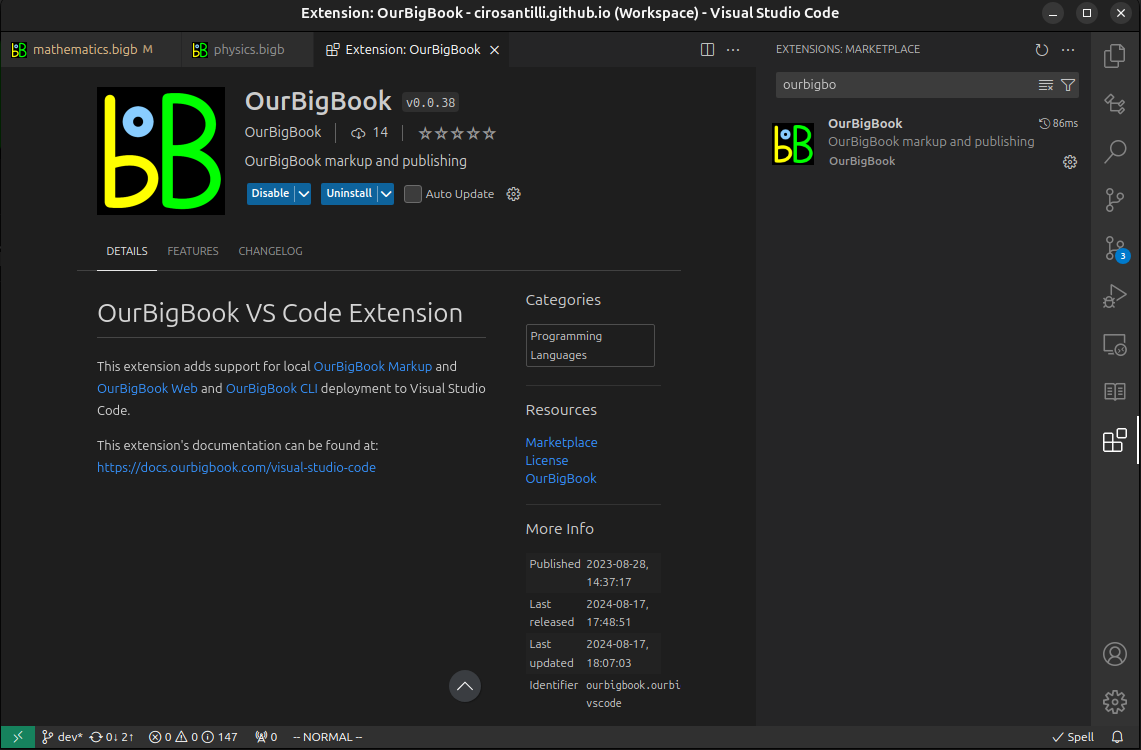
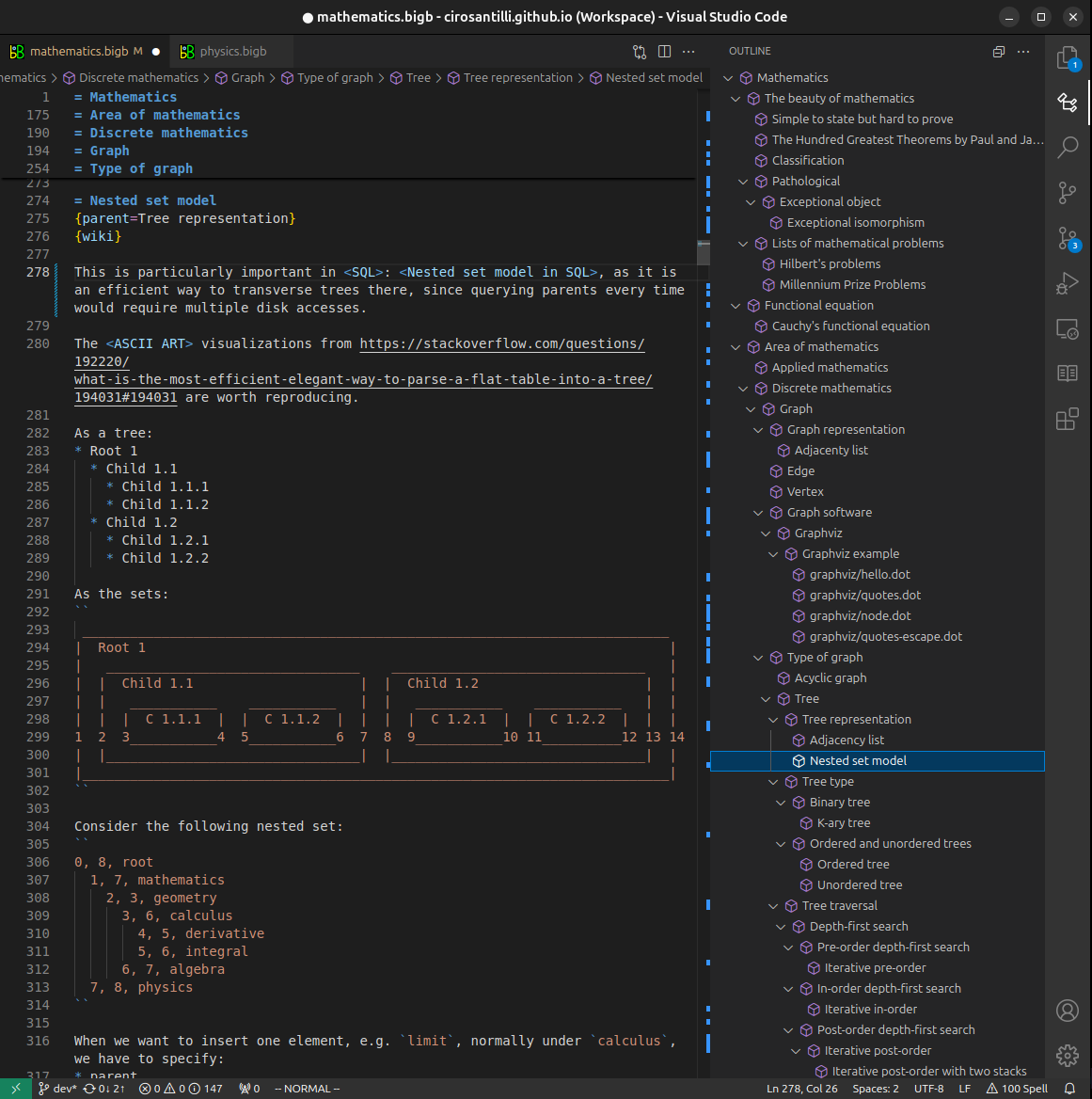
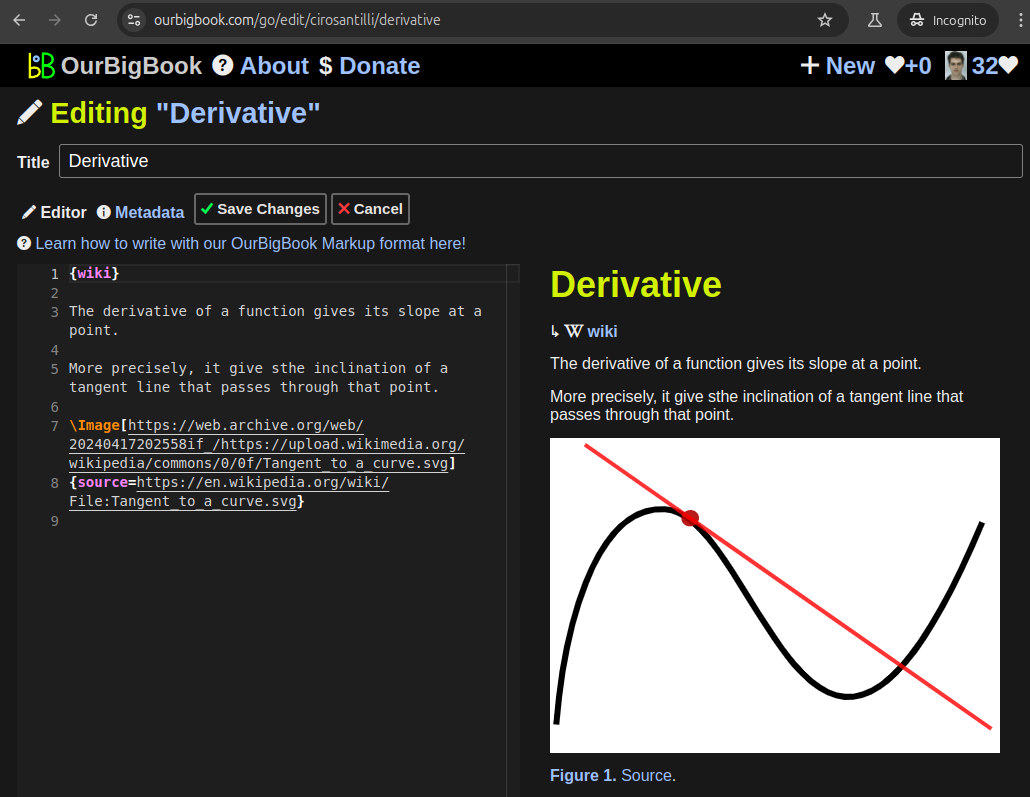
Figure 3. Visual Studio Code extension installation.Figure 4. Visual Studio Code extension tree navigation.Figure 5. Web editor. You can also edit articles on the Web editor without installing anything locally.Video 3. Edit locally and publish demo. Source. This shows editing OurBigBook Markup and publishing it using the Visual Studio Code extension.Video 4. OurBigBook Visual Studio Code extension editing and navigation demo. Source. - Infinitely deep tables of contents:
All our software is open source and hosted at: github.com/ourbigbook/ourbigbook
Further documentation can be found at: docs.ourbigbook.com
Feel free to reach our to us for any help or suggestions: docs.ourbigbook.com/#contact